Amplification-free direct detection of Ebola virus on a hybrid optofluidic platform
Low-complexity detection of infectious diseases with high sensitivity and specificity is urgently needed, especially in resource-limited settings. Optofluidic integration combines clinical sample preparation with optical sensing on a single chip-scale system, enabling the direct, amplification-free detection of single RNA from Ebola viruses. The optofluidic system fulfils all key requirements for chip-based clinical analysis, including a low limit of detection, wide dynamic range, and the ability to detect multiple pathogens simultaneously.
by Dr Hong Cai, Prof. Aaron R. Hawkins and Prof. Holger Schmidt
Introduction
The recent Ebola and Zika outbreaks [1, 2] have made it clear that viral infections continue to pose diverse and widespread threats to humanity. Resource-limited settings, in particular, call for diagnostic devices and technologies that are robust and feature relatively low complexity for easy handling by potentially unskilled personnel. At the same time, such instruments need to fulfil all the technical requirements for accurate and reliable diagnosis. These include a limit of detection and dynamic range that are compatible with clinically observed viral loads as well as the ability to carry out multiplexed differential detection by screening simultaneously for several pathogens with similar clinical symptoms.
The ‘gold standard’ test for hemorrhagic fevers as well as other infectious diseases is real-time polymerase chain reaction (RT-PCR) [3]. PCR fulfils the sensitivity and specificity requirement for clinical testing. However, it is not ideal for resource-limited environments and point-of-care applications because of to its complexity. An alternative economic and portable option is antigen-capture enzyme-linked immunosorbent assay (ELISA) testing. However, ELISA requires more highly concentrated samples and thus its clinical application, especially for early disease detection, is restricted.
For the last two decades, the lab-on-chip approach, which features a small footprint and sample volume, has been considered as a promising candidate for the next generation low-complexity medical diagnostics [4]. Among all the approaches, optofluidics, which integrates optics and microfluidics in the same platform, has received increased attention [5, 6]. Microfluidics is ideal for performing biological sample processing on a chip-scale level and leads to miniaturization and simplification of the current diagnostic system. If it can be integrated with an optical sensing/read-out platform that enables high detection sensitivity down to the single pathogen level, an analytic system for which nucleic acid amplification is no longer needed becomes possible.
In order to detect single molecular biomarkers and bioparticles, an in-flow based detection scheme is preferred. In a typical in-flow detection scheme, bioparticles are transported to the sensing region in a stream of gas or liquid where they are detected in transient fashion as they pass an optical interrogation region [7, 8]. Therefore, fast read-out of the optical signal from single bioparticles in sequence can be achieved, and many concerns associated with traditional surface-based sensing schemes such as unwanted nonspecific binding, probe photobleaching, and diffusion-limited transportation are eliminated.
Anti-resonant reflecting optical waveguides (ARROWs) have been proven to be highly efficient in detecting single bioparticles. By properly designing a Fabry–Perot reflector surrounding a hollow channel, light can propagate inside the ARROWs. Therefore, ARROWs confine both liquid and light in the same microfluidic channel, such that light and matter have near-perfect overlap and the sensing capability is maximized [8, 9]. Figure 1(a) shows a cross-section of a liquid-core ARROW using state-of-the-art fabrication technology [9]. Moreover, a two-dimensional photonic sensing platform can be constructed with lithography patterning. Figure 1(b) shows an ARROW platform with solid-core and liquid-core ARROWs crossing orthogonally. Excitation light from an external laser is confined in the solid-core ARROW, producing a few-micron-wide optical mode in the intersecting region. Liquid flow is generated inside the liquid-core ARROW to transport the bioparticles to the excitation volume which is of the order of femtolitres for typical waveguide and channel dimensions of a few micrometers. Optical read-out is extracted orthogonally through the liquid-core ARROW to achieve a low-noise signal, sufficient for reaching single particle fluorescence detection.
Besides the optical sensing aspect, miniaturizing and optimizing sample preparation is equally important in order to achieve a complete bioanalysis detection system. The ARROW-based optofluidic system is particularly well suited for such hybrid integration strategies. The planar optical layout based on intersecting solid-core and target-carrying liquid-core waveguides leaves the third dimension open for vertical integration of other functionalities. A separate microfluidic sample processing layer can be made and optimized and then connected to the ARROW platform (Fig. 1c) [10, 11]. Through this approach, we can perform multiple sample preparation steps, such as mixing, distributing, sorting and pre-concentrating on the microfluidic layer and transfer the sample to the ARROW chip for sensing without compromising each of the layers’ performance [10].
Amplification-free detection of Ebola nucleic acids on an opto-
fluidic system
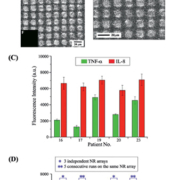
In our recent work, Zaire Ebola virus RNA detection from clinical samples has been demonstrated in a hybrid optofluidic ARROW system [12]. Through a strain-specific solid-phase extraction method, we extracted and labelled target RNA from Ebola infected Vero cells and put them through the optofluidic chip for detection. The ARROW chip provided a sequence of optical signals when individual fluorescent virus RNAs passed through the small excitation volume. Figure 1(d) shows the recorded digital RNA counts at low concentration levels of from 2.1×102 to ~2.1×104 pfu/mL within one second. We were able to detect six orders of magnitude of the clinical concentration range using the ARROW chip only. The lower concentration limit is determined by the detection time, which was set to be 10 min maximum. As a negative control, we used the same method to test for Sudan Ebola virus and Marburg virus. Our results showed no detectable signals and thus our method is target specific.
In order to incorporate critical sample processing steps and detect RNA at even lower concentrations within 10 min, we adapted a programmable microfluidic chip – an automaton – to handle processing of larger sample volumes (Fig. 1b). The polydimethylsiloxane (PDMS) based automaton chip consists of a two-layer microvalve array. Each valve’s state is controlled individually by the top pneumatic layer through a reprogrammable software program. We used the automaton chip to perform an extra pre-concentration step by processing a large amount of clinical sample. We washed, released and labelled the RNA on the same automaton chip after pre-concentration. Through ~460× concentration, the virus detection limit was improved down to 0.2 pfu/mL, with seven orders of magnitude of concentration range (Fig. 1e). This demonstration exhibits an amplification-free chip-based virus and nucleic acid analysis technique with high sensitivity and wide dynamic range, whose performance is comparable with the gold standard, more complex PCR technique.
Wavelength division multiplexing detection
ARROWs also enable simultaneous detection of multiple pathogens through the wavelength division multiplexing (WDM) technique [13]. WDM is generated using a multi-mode interferometer waveguide (MMI). When an MMI is excited by a single optical mode, all of the modes inside the MMI propagate at different phase velocities. When a constructive interference condition is satisfied, various numbers of self-imaging spots which resemble the excitation mode are formed along the MMI. This allows us to design an MMI section that intersects the fluidic channel, where multiple excitation spots are generated (Fig. 2a). As a fluorescent target flows past this excitation region, multi-peak signals are recorded in the time domain. For a single wavelength excitation, the fixed pattern multi-peak detection enables a signal-to-noise improvement compared to single-mode detection [14].
For a given MMI, the number of the self-imaging spots is wavelength dependent. We can generate various spot patterns at various laser wavelengths. For example, 9, 8 and 7 excitation spots are generated using 488nm, 553nm and 633nm lasers (Fig. 2b). With this approach, multiple targets labelled with different dye can be distinguished by the number and spacing of the peaks in the detected signal. Figure 2(c) shows influenza virus H1N1 and H3N2, which were labelled with different dye, generating 9 and 6 peaks in the time domain, respectively. We also labelled H2N2 virus with a combination of these two dyes which resulted in a superposition of the 9-spot and 6-spot fluorescence signals (Fig. 2c, bottom). A signal-processing algorithm checks for the presence of signals at the two characteristic time delays and can easily identify the mixed-labelled virus particle. This technique was shown to discriminate between three influenza subtypes, again with single virus sensitivity, using only two excitation colours. Thus, the ARROW-based platform has now met all the fundamental requirements for clinical virus detection using single particle sensing.
Conclusion
For the next generation of medical diagnostic devices, low-complexity detection with high sensitivity and specificity is required on the detection side, along with small footprint and multi-functional analyte handling on the sample processing side. In-flow based optofluidic devices in which both analyte handling and optical sensing are carried out on the chip scale are promising candidates. Using our ARROW-based optofluidic system, we demonstrated multi-stage sample processing and detection of clinical Zaire Ebola virus samples using hybrid integration. We also demonstrated wavelength multiplex detection of multiple analytes at the same time. This fulfils all quantitative requirements for clinical virus detection. Therefore, a fully integrated microsystem for front-to-back amplification-free virus analysis is within reach.
References
1. Fact sheet. The top 10 causes of death. World Health Organization 2014. (http://www.who.int/mediacentre/factsheets/fs310/en/).
2. Fact sheet. Zika virus. World Health Organization 2016. (http://www.who.int/mediacentre/factsheets/zika/en/).
3. Kuypers J, Wright N, Morrow R. Evaluation of quantitative and type-specific real-time RT-PCR assays for detection of respiratory syncytial virus in respiratory specimens from children. J Clin Virol. 2004; 31: 123–129.
4. Craighead H. Future lab-on-a-chip technologies for interrogating individual molecules. Nature 2006; 442: 387–393.
5. Fan X, White IM. Optofluidic microsystems for chemical and biological analysis. Nature Photon. 2011; 5: 591–607.
6. Schmidt H, Hawkins AR. The photonic integration of non-solid media using optofluidics. Nature Photon. 2011; 5: 598–604.
7. Zhu H, White IM, Suter JD, Zourob M, Fan X. Opto-fluidic micro-ring resonator for sensitive label-free viral detection. Analyst 2008; 133: 356–360.
8. Bernini R, Campopiano S, Zeni L, Sarro PM. ARROW optical waveguides based sensors. Sensors and Actuators B 2004; 100: 143–146.
9. Yin D, Barber JP, Hawkins AR, Deamer DW, Schmidt H. Integrated optical waveguides with liquid cores. Appl Phys Lett. 2004; 85: 3477–3479.
10. Parks JW, Cai H, Zempoaltecatl L, Yuzvinsky TD, Leake K, Hawkins AR, Schmidt H. Hybrid optofluidic integration. Lab Chip 2013; 13: 4118–4123.
11. Testa G, Persichetti G, Sarro, PM, Bernini R. A hybrid silicon-PDMS optofluidic platform for sensing applications. Biomed Opt Express 2014; 5: 417–426.
12. Cai H, Parks JW, Wall TA, Stott MA, Stambaugh A, Alfson K, Griffiths A, Mathies RA, Carrion R, Patterson JL, Hawkins AR, Schmidt H. Optofluidic analysis system for amplification-free, direct detection of Ebola infection. Scientific Reports 2015; 5: 14494.
13. Ozcelik D, Parks JW, Wall TA, Stott MA, Cai H, Parks JW, Hawkins AR, Schmidt H. Optofluidic wavelength division multiplexing for single-virus detection. Proc Nat Acad Sci U S A 2015; 112: 12933–12937.
14. Ozcelik D, Stott MA, Parks JW, Black JA, Wall TA, Hawkins AR, Schmidt H. Signal-to-noise enhancement in optical detection of single viruses with multi-spot excitation, IEEE J Sel Top Quant Elec. 2016; DOI: 10.1109/JSTQE.2015.2503321.
The authors
Hong Cai1 PhD, Aaron R. Hawkins2 PhD, Holger Schmidt*1 PhD
1School of Engineering, University of California Santa Cruz, Street, Santa Cruz, CA 95064 USA
2ECEn Department, 459 Clyde Building, Brigham Young University, Provo, UT 84602 USA
*Corresponding author
E-mail: hschmidt@soe.ucsc.edu