Biomarker panels for the diagnosis of sepsis
Sepsis is a complex syndrome associated with significant morbidity and mortality. If detected and treated early, septic patients have better prognoses. Unfortunately, identification of sepsis is challenging because its pathophysiology is complex and its clinical signs and symptoms overlap with other inflammatory diseases. This review discusses emerging biomarker panels and their ability to predict sepsis in critically ill patients.
by Dr A. Woodworth and Dr J. Colon-Franco
Sepsis and SIRS: International definitions
Definitions of sepsis and related conditions date back to the 1991 consensus conference held by the American College of Chest Physicians and the Society of Critical Care Medicine [1].
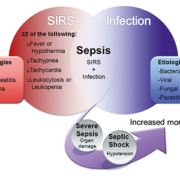
This consensus group introduced the term SIRS to describe the systemic inflammatory response syndrome, a normal response to infection and non-infectious insults like trauma, pancreatitis, and burns [Figure 1]. In SIRS two or more of the following clinical signs manifest: abnormal body temperature (fever or hypothermia), tachypnea, tachycardia and abnormal white blood cell count (leukocytosis or leukopenia).
The consensus group defined sepsis as the presence of SIRS along with a documented infection. Left untreated, septic patients develop severe sepsis, characterized by organ dysfunction, and ultimately septic shock, characterized by organ failure, hypotension and decreased peripheral perfusion. Revision of these definitions in 2001, added ‘suspected infection’ to the classification of sepsis to address numerous clinical cases where microorganisms cannot be confirmed.
For over 20 years, these definitions have provided uniformity in clinical disease recognition and better characterized patient populations for sepsis research. Although recognition of SIRS is relatively straightforward, identification of patients with sepsis among those with SIRS remains challenging. This is due, in part, to overlapping clinical signs and symptoms between SIRS and sepsis as well as inherent difficulty in confirming infectious causes of SIRS.
Sepsis and SIRS: Pathobiology and related syndromes
Sepsis pathobiology is complex and not well characterized [Figure 2] [1]. Historical models describing an overactive proinflammatory response to infection likely oversimplify the process. Sepsis experts now support two distinct pathogenesis models for the progression of sepsis. The sequential response model describes an initial proinflammatory response to a pathogen, SIRS, followed by a compensatory anti-inflammatory response syndrome (CARS). In the second model, known as the mixed antagonist response syndrome, SIRS and CARS occur simultaneously and achieve homeostasis. Severe sepsis and septic shock are associated with an imbalance in the SIRS/CARS equilibrium. Although recent research supports the second model [2], larger studies exploring the underlying pathobiology, including expression of pro- and anti-inflammatory molecules throughout the course of sepsis pathogenesis are needed.
Sepsis diagnosis
Rapid diagnosis and treatment of sepsis reduces mortality. The ‘gold standard’ for sepsis diagnosis is identification of an infectious microorganism in patients with SIRS. Traditionally, pathogens in blood, urine or other body fluids were detected in the laboratory by culturing. Unfortunately, cultures have limited utility because some pathogens are slow growing and contamination is common, leading to a high number of false negative and positive results. Despite their disadvantages, identification of the infecting agent as well as its antibiotic susceptibility and resistance patterns remains crucial to administer or adjust antimicrobial treatment. Direct identification of a pathogen through molecular and proteomic-based approaches may help overcome these disadvantages [3].
Because of its non-specific clinical symptoms and the limited utility of bacterial cultures, researchers have looked to biomarkers to diagnose sepsis. Currently, the diagnostic utility of sepsis biomarkers is limited to confirming, ruling out sepsis or stratifying patients based on disease severity. Lactate, C-reactive protein (CRP) and procalcitonin (PCT) assist in the work-up of patients with suspected sepsis. Lactate, the end product of anaerobic glycolysis, is increased in septic shock and other conditions as a result of excessive energy demand, tissue hypoxia, and/or impaired metabolic pathways.
The Surviving Sepsis Campaign, an international collaboration developed to improve the management, diagnosis, and treatment and reduce mortality rates of sepsis, advocates measuring blood lactate within 6 h of presentation in patients with suspected sepsis [1]. A lactate concentration >4 mmol/L (36 mg/dL) is associated with increased morbidity and mortality and is used to guide sepsis resuscitation protocols. Lactate concentrations increase with severity of sepsis and are most useful for diagnosing septic shock, but lack diagnostic strength to discriminate early sepsis from SIRS.
Expression of proinflammatory molecules is markedly up-regulated in early sepsis. CRP and PCT expression is stimulated by proinflammatory cytokines. CRP is an acute phase reactant that is up-regulated in inflammatory processes, and is not specific for sepsis. PCT, the precursor of calcitonin in thyroidal C-cells, is systemically produced in non-thyroidal tissue in response to inflammation and infection. Compared to CRP, PCT more accurately distinguishes SIRS from sepsis [1, 4]. In critically ill adults, the diagnostic strength of PCT to distinguish sepsis from SIRS is low [1, 4]. This may be due to the fact that like CRP, PCT is overexpressed in non-infectious inflammatory states like surgery or trauma. Unlike CRP, PCT concentrations correlate with sepsis severity. Both CRP and PCT can predict prognosis and response to therapy in septic patients. PCT is also useful in ruling out bacterial infections and is used in algorithms guiding antimicrobial therapy in critically-ill patients [4]. However, because of its questionable diagnostic utility, PCT testing is not universally used in clinical practice.
Thousands of studies have investigated the clinical and diagnostic utility of hundreds of sepsis biomarkers. A recent review of relevant clinical and experimental studies identified 178 proposed sepsis biomarkers [4]. Besides PCT and CRP, 34 others were investigated as diagnostic markers for sepsis. None had sufficient diagnostic strength to differentiate septic patients from those with non-infectious SIRS.
Multiple biomarker panels for sepsis diagnosis
As our understanding of the underlying mechanisms of sepsis evolved, it became evident that a single biomarker could not identify all patients with this heterogeneous syndrome. Instead, a panel of biomarkers, consisting of molecules secreted in the blood throughout the disease process, may better predict sepsis among patients with systemic inflammation [3].
Recent studies [Table 1] explored the utility of novel multimarker panels to predict sepsis. In a prospective cohort study of 151 emergency department (ED) patients with SIRS, both panels of 3 and 6 biomarkers showed superior diagnostic utility for detection of bacterial infection compared to any single biomarker [Table 1; Study #1] [5]. In a separate cohort of 342 ED patients with SIRS, adding 1 to 3 biomarkers and/or clinical parameters did not improve upon PCT alone to predict bacteremia [Table 1; #2] [6]. In the latter study, only patients with documented positive blood cultures were included, excluding possible infection in other sites or fluids and patients with false negative cultures.
In a retrospective pilot study at our institution, 10 inflammatory biomarkers, chosen because of their expression pattern during SIRS and/or CARS, were measured in 63 critically-ill patients with SIRS [Table 1; #3]. Panels of 2 to 6 inflammatory biomarkers measured in multiplex were better able to identify sepsis among patients with SIRS compared to single markers. Because of the small sample size, a 2-marker panel was most predictive of sepsis. PCT and CRP showed limited diagnostic utility alone or in combination with other biomarkers [7]. In a second retrospective cohort study we evaluated the diagnostic utility of 5 inflammatory biomarkers up-regulated in SIRS and/or CARS in 169 ICU patients with SIRS [Table 1; #4]. The 5-biomarker panel outperformed any single biomarker to predict sepsis on the day that patients developed SIRS [8]. Studies are ongoing to validate these findings in a larger population and to compare these results with the diagnostic performance of PCT.
A sepsis risk score, generated from results of multiple biomarkers, may allow easy adoption of these panels into clinical practice [3, 9]. A study evaluating the plasma concentrations of 5 proinflammatory molecules demonstrated that, compared to individual markers, a sepsis score consisting of at least 2 biomarkers elevated above their respective cut-offs better discriminated between SIRS and sepsis in ICU patients [Table 1; #5] [10]. Gibot and colleagues investigated the concentration of three biomarkers in 300 patients consecutively admitted into the ICU [Table 1; #6] [9]. A bioscore of 0, 1, 2 or 3 was assigned based on the number of positive biomarkers (above pre-defined cut-off values). The bioscore surpassed the diagnostic strength of any of the individual biomarker results for the prediction of sepsis. This model was validated in a separate cohort of 228 patients presenting with clinical signs of sepsis. In these studies, the sepsis score strategy was practical with superior diagnostic utility.
Conclusions
Early identification and treatment of septic patients reduces mortality, however, signs and symptoms of early sepsis are similar to non-infectious SIRS. To date, no single biomarker has ample diagnostic strength to identify septic patients among a critically-ill population. A panel of biomarkers may better distinguish patients with sepsis from those with non-infectious SIRS. Most findings are from preliminary studies with small patient cohorts and require additional validation studies. These should be conducted in larger, multicentre populations with distinct validation cohorts. Rapid, automated, multiplexing platforms and/or point-of-care technologies may be necessary to obtain timely results for these multimarker sepsis panels. Combining biomarkers into equations or sepsis-scores that yield an interpretable and meaningful result is paramount for their clinical adoption.
In conclusion, using combinations of biomarkers to predict sepsis is an attractive strategy that may improve real time assessments and reduce morbidity and mortality in septic patients.
References
1. Faix JD. Established and novel biomarkers of sepsis. Biomark Med 2011; 5(2): 117–130.
2. Osuchowski MF, et al. Sepsis chronically in MARS: systemic cytokine responses are always mixed regardless of the outcome, magnitude, or phase of sepsis. J Immunol 2012; 189(9): 4648–56.
3. Casserly B, Read R, Levy MM. Multimarker panels in sepsis. Crit Care Clin 2011; 27(2): 391–405.
4. Pierrakos C, Vincent JL. Sepsis biomarkers: a review. Crit Care 2010; 14(1): R15.
5. Kofoed K, et al. Use of plasma C-reactive protein, procalcitonin, neutrophils, macrophage migration inhibitory factor, soluble urokinase-type plasminogen activator receptor, and soluble triggering receptor expressed on myeloid cells-1 in combination to diagnose infections: a prospective study. Crit Care 2007; 11(2): R38.
6. Tromp M, et al. Serial and panel analyses of biomarkers do not improve the prediction of bacteremia compared to one procalcitonin measurement. J Infect 2012; 65(4): 292–301.
7. Pyle AL, et al. Multiplex cytokine analysis for the differentiation of SIRS and sepsis. Am J Clin Pathol 2010; 134: 509.
8. Pyle AL, et al. A multi-marker approach to differentiate sepsis from SIRS. Am J Clin Pathol 2011; 136: 468–469.
9. Gibot S, et al. Combination biomarkers to diagnose sepsis in the critically ill patient. Am J Respir Crit Care Med 2012; 186(1): 65–71.
10. Selberg O, et al. Discrimination of sepsis and systemic inflammatory response syndrome by determination of circulating plasma concentrations of procalcitonin, protein complement 3a, and interleukin-6. Crit Care Med 2000; 28(8): 2793–2798.
The authors
Alison Woodworth, PhD, DABCC, FACB
Assistant Professor, Pathology, Microbiology and Immunology
Director, Esoteric Chemistry
Vanderbilt University Medical Center
Nashville, TN, USA
Jessica M. Colón-Franco, PhD
Clinical Chemistry Fellow
Department of Pathology, Microbiology and Immunology
Vanderbilt University Medical Center
Nashville, TN, USA
E-mail:
Alison.Woodworth@Vanderbilt.Edu