Liquid chromatography-tandem mass spectrometry: an introduction
The use of liquid chromatography-tandem mass spectrometry for clinical analysis is on the increase. This article describes what it is, why it can offer significant improvements over traditional assays and the limitations to be aware of.
by Dr N. Homer
Introduction
Clinical biochemistry laboratories frequently use radioimmunoassays (RIA) and enzyme-linked immunosorbent assays (ELISA) for analysis of blood and urine. However, these techniques are plagued by issues of cross-reactivity and are only suited to look at one analyte at a time [1]. The use of mass spectrometry (MS) techniques has increased since 2007 when the American Endocrine Society recognized the importance of tandem MS and issued a statement recommending the use of liquid chromatography-tandem mass spectrometry (LC-MS/MS) for the determination of endogenous steroid hormones over more traditional technologies such as immunoassays [2]. This has led to the widespread adoption of LC-MS/MS in clinical biochemistry laboratories, in direct response to this recommendation.
What is liquid chromatography-mass spectrometry?
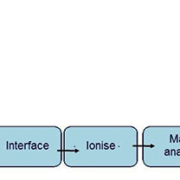
Mass spectrometry is a technique that measures charged molecules or ions in the gaseous state. Samples are introduced into an ion source, ionized and then separated in a mass analyser according to their mass-to-charge ratio (m/z) and then characterized by their relative abundances. Coupled to chromatographic separation techniques such as gas chromatography (GC) or liquid chromatography (LC), MS is considered to be the ‘gold standard’ for validation of quantitative analytical assays. An overview of how a typical chromatograph-mass spectrometer is set up is shown in Figure 1.
Following separation by a chromatography system the sample is introduced into an ion source at the front end of the mass spectrometer. Ionization modes include atmospheric pressure ionization (API), such as electrospray ionization (ESI) and atmospheric pressure chemical ionization (APCI), and matrix-assisted laser desorption/ionization (MALDI). ESI is most typically used to ionize the biomolecules encountered in clinical samples.
Once ionized the mass analyser separates the ions according to their m/z. Mass analysers include magnetic or electric sectors, time-of-flight (ToF) tubes, quadrupoles and two-dimensional and three-dimensional ion traps. Softer ionization techniques, which generally leave the molecule intact, such as ESI, have led to the use of quadrupole mass analysers. These consist of four parallel rods or poles, generally of hyperbolic cross-section, through which ions are passed and separated.
Tandem mass spectrometry (often termed MS/MS) technology increases the specificity of MS significantly. There are a number of modes that a tandem mass spectrometer can be operated under, depending on the requirement of the experiment (Fig. 2). Tandem MS requires two or more mass analysers to be placed in sequence and the ions are fragmented in a collision cell to give structural information. Trace analysis of complex biological matrices is ideally suited to tandem MS instruments, operated in selected reaction monitoring (SRM) mode. In addition, linear ion traps as the third mass analyser are also increasing in popularity as they offer additional structural identification and specificity.
Sample preparation and liquid chromatography method development
Clinical samples are complex biological matrices and contain interferences that can lead to so-called matrix effects within the mass spectrometer. For validated assays, samples are prepared by addition of an internal standard followed by extraction to remove as much of the interferences as possible. The internal standard is either a closely related analogue of the compound of interest, or a stable isotope labelled version of the compound, enriched with at least two atoms of 13C, 2H or 15N.
Sample preparation methods commonly applied to clinical samples include protein precipitation with an organic solvent, liquid–liquid extraction (LLE) or solid-phase extraction (SPE). If sample clean-up is not sufficient it can lead to matrix effects, including ion suppression of the analyte, usually observed as a loss of response. This affects the detection limits, accuracy and precision of the assay. Various ion suppression tests have been developed and these are an important part of the method validation set-up required for clinical MS assays. The two most effective ways of avoiding ion suppression are improved sample extraction and optimized chromatographic selectivity.
On-line multidimensional chromatography technology allows an unextracted sample to be introduced into the chromatography apparatus and can lead to faster analysis. These systems generally consist of multi-channel switching valves, on-line SPE cartridges and analytical columns ahead of ESI-LC-MS/MS. Steroids and isoprostanes are often analysed in this manner [3, 4].
Liquid chromatography
Once prepared, a sample is introduced into the LC system which consists of a pump and an analytical column. The purpose of the chromatography system is to separate the components of the sample as much as possible, before introduction into the mass spectrometer. Analytical LC columns are stainless steel tubes that are packed with tiny silica beads. The type of LC used in clinical analysis is usually reversed-phase chromatography as the silica beads are generally chemically modified. Typically, samples are introduced onto the column in a highly aqueous phase, the analytes associate with the chemically-modified packed silica beads and are washed off the column with a high organic solvent such as methanol or acetonitrile.
Once the ionization and mass spectrometer parameters have been optimized, much of the method development falls to the chromatography and the importance of this stage should not be underestimated. It is imperative that co-eluting compounds do not interfere with the analytical peaks of interest. In recent years there has been a trend for fast analysis in LC-MS/MS; however, this does not always give a robust assay. In addition, it is important to be aware of isobaric compounds (same mass) and [M+2] isotopomers. An example of isotopomers is that of the stress hormone cortisol (m/z 363) and its inactive form cortisone (m/z 361). In an LC-MS/MS assay for cortisol it is essential to have two separate peaks for cortisol and cortisone, otherwise the risk of isotopomers of cortisone contributing to cortisol would lead to an over-estimation of cortisol in the sample (Fig. 3).
There are a number of parameters that can be altered in LC and these in turn alter the selectivity of the column, that is the order and rate at which the components elute. Parameters that can be influenced include column temperature, mobile phase pH, composition and flow rate, column dimensions, column particle size and the nature of course the chemical modification of the particles too.
Analytical LC column technology is continuously improving. The better the resolution, which is simply how well separated each peak is, the better the assay. Sub-2 µm particles have been introduced in the past decade, which generate sharp peaks and excellent resolution with improved capacity over the more traditional 3–5 µm particles. However, the smaller particle size leads to high backpressure and requires specific LC pumps that can withstand these ultra-high pressures (UHPLC, ultra-high performance LC). To reduce the need for new instrumentation, LC columns packed with fused-core particles ~2.5 µm have been developed to allow separation comparable to sub-2 µm particles. The backpressure generated by these fused-core particles is significantly less than the sub-2 µm particle columns and exclude the requirement of high-pressure capable LC pumps and fittings.
Considerations when establishing an LC-MS/MS clinical biochemistry method
As with all techniques, there are drawbacks to LC-MS/MS. The instrumentation and software can be complex and requires regular maintenance, although manufacturers are addressing this perception by introducing simpler software interfaces with dedicated instrument support for method development and even fool-proof methods, guaranteed by the provider. Also, some compounds are not amenable to ionization due to their chemical nature, but chemical modification before analysis can improve the chance of ionization efficiency, so all is not lost.
Summary
The benefits that MS offers over other traditional assay techniques have seen an increase in the number of assays using this methodology. The analysis of steroid hormones by MS is a well-documented area. Other commonly encountered uses include newborn screening for congenital metabolic diseases such as aminoacidopathies and fatty acid oxidation disorders, multi-analyte therapeutic drug monitoring, oncology drugs, anti-virals, toxicants and drugs of abuse screening and analysis of endogenous peptides [3, 4, 5].
One area that is continuing to gain interest in clinical research is high-resolution MS (HRMS) [5]. This allows for accurate mass determination over a defined mass range, which differs from the targeted analysis approach used by triple quadrupole MS. With technological improvement in the linear range of HRMS instruments to match that of triple quadrupoles, it seems likely that the benefits of HRMS will also be exploited by the clinical biochemistry field, in addition to LC-MS/MS analysis.
The range of clinical applications of MS outlined is broad and constantly expanding. Much research is being conducted in the pioneering fields of proteomics and metabolomics. In recent years the emergence of imaging mass spectrometry also offers exciting possibilities for the future and there is no doubt that MS will continue to feature heavily in the clinical biochemistry laboratory and function as an important clinical research tool.
References
1. Penning TM, et al. Liquid chromatography-mass spectrometry (LC-MS) of steroid hormone metabolites and its applications. J Ster Biochem Mol Biol. 2010; 121: 546–555.
2. Rosner W, et al. Position statement: utility, limitations, and pitfalls in measuring testosterone: an Endocrine Society position statement. J Clin Endocrinol Metab. 2007; 92: 405–413.
3. Shushan B. A review of clinical diagnostic applications of liquid chromatography-tandem mass spectrometry. Mass Spectrom Rev. 2010; 29: 930–944.
4. Chace DH, et al. Use of tandem mass spectrometry for multianalyte screening of dried blood specimens from newborns. Clin Chem. 2003; 49: 1797–1817.
5. Jiwan J-LH, et al. HPLC-high resolution mass spectrometry in clinical laboratory? Clin Biochem. 2011; 44: 136–147.
The author
Natalie Homer PhD
CRF Mass Spectrometry Core Laboratory, Queen’s Medical Research Institute, University of Edinburgh
E-mail: n.z.m.homer@ed.ac.uk