Meeting clinical needs with high performance viral load assays, workflow improvements and reduced turnaround times
Niguarda Hospital in Milan is one of Italy’s leading general hospitals, and provides an extensive range of medical disciplines for adults and children throughout the Lombardy region and beyond.
Our hospital’s Department of Laboratory Medicine aim is to offer a complete, continuous and prompt diagnostic laboratory testing service, in order to guarantee effective support for this widespread clinical demand, and is committed to research into automation and analysis to ensure this is maintained. Our busy Molecular Biology Laboratory performed an estimated 40,000 tests in 2015, which is approximately 10% increase on the previous year.
The growing annual molecular workload is attributed, in part, to the development of new therapeutic strategies. Our staff, consisting of 8 laboratory technicians, one director and one manager, work 5 days per week and are expected to cope with increased workloads and demands for reduced turnaround times without any increase in resources, in terms of the number of staff and costs.
A large proportion of the molecular biology workload consists of viral load measurements for human immunodeficiency virus type 1 (HIV-1), hepatitis C virus (HCV), hepatitis B virus (HBV) and cytomegalovirus (CMV) (figure 1).
With a very important Italian transplant centre located at Niguarda Hospital, CMV analyses are vital and results are needed quickly, without delay. In addition, the laboratory performs viral load measurements for HIV-1, HCV, and HBV in order to evaluate and monitor therapeutic response. In these instances, rapid results are extremely important for patient management decisions, for example to maintain or change treatment.
Since 2005, these measurements have been performed using our laboratory’s current method, which has separate sample preparation and amplification/detection platforms. These are situated on separate benches within the same room, with one sample preparation system in another room. The accuracy and precision of this method is good, however, in order to be cost effective, it is necessary to optimize the size of the batches. Since they can’t be processed in the same day, sample test tubes often need to be collected and stored for several days, which increases the turnaround time considerably. In addition, this method involves many manual steps, which demand time, space and coordination of work between different members of staff.
A new automated molecular diagnostics method
As part of our Department of Laboratory Medicine’s investigations into increased automation in the laboratory, Niguarda Hospital became a beta trial site for the new DxN VERIS Molecular Diagnostics System (Beckman Coulter), which consolidates DNA extraction, nucleic acid amplification, quantification and detection onto a single automated instrument for a number of molecular targets, including HIV-1, HCV, HBV and CMV.
The first step in assessing the DxN VERIS was to validate the assays in order to determine whether their performance is comparable with our laboratory’s existing method. Daily quality control measurements demonstrated good performance of the VERIS HBV assay for high level, low level and negative HBV samples (table 1). This assay was also shown to have excellent linearity within the range of 1.68 – 8.82 Log IU/mL, a limit of detection of 6.82 IU/mL, and good precision, achieving within run and between run mean standard deviations of less than 0.16 (table 2).
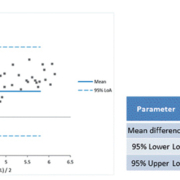
A series of performance evaluation studies, conducted in several laboratories around the world, have demonstrated that the VERIS HBV, HCV, HIV-1 and CMV assays have comparable precision, sensitivity and linearity to a range of alternative, commercially available viral load methods [1-13]. In accordance with these findings, the VERIS HBV assay correlated well with the existing method at Niguarda Hospital (Abbott m2000) and, indeed, detected HBV DNA in 23 samples that were negative using the current method, 22 of which were found to be positive by one or more serology assay (table 3). Regarding the 55 specimens that were quantified both with DxN VERIS and Abbott m2000, 7 of them had an HBV DNA concentration discordant for more than 1 Log.
Comparable performance, including sensitivity and specificity, was achieved for each of the DxN VERIS assays: HIV-1, HCV, HBV and CMV.
Workflow improvements
In addition to validating the performance of the VERIS assays, a time/workflow analysis study was performed at Niguarda Hospital by Nexus Global Solutions (Plano, Texas, USA). The study compared workflows and time to results between the current viral load method for HIV-1, HCV, HBV and CMV (Abbott m2000sp and m2000rt systems) and the new DxN VERIS Molecular Diagnostic System.
By reducing manual intervention and automating processes from sample loading to reporting of results, the DxN VERIS offers the potential to transform clinical laboratory workflows. Each assay is supplied in a unique, single cartridge system, and all consumables and reagents are stored on board the system, which cuts preparation time compared to alternative methods. In addition, unlike traditional plate-based systems, there is no need to batch assays. The DxN VERIS allows true, single sample random access, which means that viral load assays can be performed as soon as they arrive in the laboratory. This, combined with short assay runtimes, ensures rapid turnaround of results and, since there are no empty plate wells, wastage and consumable costs are reduced.
The comparative time/workflow analysis in our study revealed that DxN VERIS involved only 10 steps and required just five reagents, compared to 26 steps and over 20 consumables for the current method, and required much less hands-on time for each of the viral load assays (figure 2). Notably, by consolidating the assay menu, time savings of up to 2 hours could be achieved.
In addition to an increase in productivity (achieving more results in an 8-hour working day), the time to the first result for the DxN VERIS was greatly reduced compared to the current method, with subsequent results available every 2.5 minutes. This is in contrast to the current method, where results are not available until the end of the assay run (table 4).
With these time savings, and by eliminating the need to batch samples, the DxN VERIS allowed much faster turnaround of results in a normal working week, with all results being reported within 8 hours of receipt, unlike the current method, which often required several days (figure 3).
The true single sample random access capability of the DxN VERIS has the potential to simplify sample management in the laboratory and to make the organization of viral load assays more fluid. It increases productivity by allowing the continuous loading of samples for different assays, eliminating the need for batching and reducing turnaround times. This is the most important advantage of random access testing for us because it increases the availability of medical reports to the different departments and is a great benefit to patient management and care by allowing more timely clinical decisions.
The DxN VERIS is easy to use with its few consumables, reduced maintenance requirements, complete automation and intuitive computer interface. By improving laboratory organization and workflows and reducing manual intervention, viral loads (which account for about 50% of the molecular workload) could be completed in a single day using the DxN VERIS. Requiring fewer people to be dedicated to this purpose, this makes it possible to accomplish more work with the same number of staff.
For further information about the DxN VERIS Molecular Diagnostic System and the VERIS assays currently available, please contact: Tiffany Page, Senior Pan European Marketing Manager Molecular Diagnostics, Email: info@beckmanmolecular.com or visit www.beckmancoulter.com/moleculardiagnostics
References
1. Williams, JA, Rodriguez, J, Wang, Z et al (2014) Poster presentation, ESCV, Prague.
2. Drago, M, Franchetti, E, Fanti, D and Gesu, GP (2015) Poster presentation, EuroMedLab, Paris.
3. Zurita, S, Gutiérrez, F, Folgueira, MD et al (2015) Poster presentation, EuroMedLab, Paris.
4. Christenson, R, Maggert, K, Ruiz, RM et al (2015) Poster presentation, ECCMID, Copenhagen.
5. Trimoulet, P, Tauzin, B, Belloc, E et al (2015) Poster presentation, EuroMedLab, Paris.
6. Gilfillan, R, Wang, Z, Xu, Y et al (2014) Poster presentation, ECCMID, Barcelona.
7. Xu, Y, Gilfillan, R, Wang, Z et al (2014) Poster presentation, ESCV, Prague.
8. Mengelle, C, Sauné, K, Haslé, C et al (2014) Poster presentation, RICAI.
9. Mengelle, C, Sauné, K, Haslé, C et al (2015) Poster presentation, ECCMID, Copenhagen.
10. Silvestro, A, Duan, H, Lim, S et al (2014) Poster presentation, ECCMID, Barcelona.
11. Li, Q, Williams, J, Maggert, K et al (2014) Poster presentation, ECCMID, Barcelona.
12. Xu, Y, Dineen, S, Annese, V et al (2014) Poster presentation, ESCV, Prague.
13. Williams, JA, Rodriguez, J, Wang, Z et al (2014) Poster presentation, ECCMID, Barcelona.
The author
Diana Fanti, Molecular Biology Laboratory Manager
Department of Laboratory Medicine, Niguarda Hospital,
Milan, Italy