Point-of-care molecular test for Zika infection
The recent emergence of the Zika virus (ZIKV) pandemic has underscored the need for a point-of-care (POC) test that can detect viral infections, fostering widespread, accurate and timely diagnostics [1, 2]. Zika infection is often asymptomatic or has comparatively mild symptoms, such as fever and chills that are common to many other infections. However, as is widely known, Zika infection during pregnancy entails substantial risks for the newborn, including severe birth defects [3]. Hence, it is highly desirable to screen women of child-bearing age and their partners for exposure to the ZIKV. Zika episodes have created bottlenecks in conventional laboratory-based diagnostic.
Zika is spread primarily by mosquitos, but sexual and perinatal transmission, as well as transmission via blood transfusions have also been reported. Specific applications for Zika diagnostics thus include determining whether infection has occurred over the course of a pregnancy, whether sexual partners harbour infection, safeguarding the blood supply and tracking the geographic range of Zika infections [4]. In order to inform investment decisions on prevention, control and response, Lee et al. [5] modelled the economic burden for various scenarios of Zika emergence across six US states, estimating total costs (direct medical, Medicaid, productivity losses) ranging from 0.5 to 2 billion US dollars.
Ideally, a POC technology will enable rapid diagnostics tests that can be performed outside of centralized laboratories such as, for example, in doctors or dentists offices, pharmacies, rural clinics, school infirmaries, border crossings or, ultimately, as an over-the-counter test for home use. Thus, the availability of a mass-produced, inexpensive POC diagnostics device with easy-to-interpret test results, and that could be used in any almost any locale by non-specialists with minimal training, would greatly expand capabilities and options for screening, surveillance, diagnostics, monitoring and therapy.
Current diagnostics technology
According to the World Health Organization (WHO), the pipeline diagnostic ZIKV kits can be categorized into: (i) antibody/antigen-based immunoassay and (ii) nucleic acid-based molecular diagnostics [6]. Immunoassay (antibody detection) for ZIKV infection utilizes envelope proteins and NS1 as targets. The major challenge is that these antibodies cross-react with other highly homologous flaviviruses such as dengue, resulting in non-specific test results [7]. IgM and IgG antibodies, typically emerge, respectively, ~4 and ~10 days after infection, but are usually undetectable until >7–14 days post-infection. Moreover, antibody responses during pregnancy may differ from those in non-pregnant individuals [8], which may adversely impact the effectiveness of immunoassay tests. Moreover, antibody tests may not discriminate between recent and historic exposure. The Food and Drug Administration (FDA) recently authorized for emergency use of the IgM Antibody Capture Enzyme-Linked Immunosorbent Assay (Zika MAC-ELISA) to detect ZIKV [8]. However, this assay requires a lab-format with delays in generating results given current demand. More importantly, these assays are a readout for exposure to ZIKV, whereas active virus infection is not determined. Currently, several companies are developing lateral flow-based rapid diagnostic test for ZIKV antibody detection [9, 10].
Molecular diagnostics-based on reverse-transcription (RT)-PCR is highly specific and sensitive, and considered the gold standard for ZIKV detection. RT-PCR is effective in serum, semen, and saliva within 14 days post-infection, and possibly much longer in urine and semen [11, 12]. Importantly, a recent study has demonstrated that ZIKV is detectable in pregnant women throughout their pregnancy [3]. Indeed, the FDA has authorized the use of the Trioplex rRT-PCR laboratory test to detect ZIKV, dengue virus, and chikungunya virus RNA, under an Emergency Use Authorization (EUA) [13]. Several research groups and companies are developing multiplexed molecular assays to concurrently detect various members of the genus Flavivirus. Most of these RT-PCR kits require, however, instrumentation and are for central laboratory use only.
Instrument-free point-of-care molecular detection of ZIKV
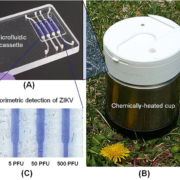
To develop inexpensive molecular detection of ZIKV without complex instrumentation, we utilized reverse-transcription loop-mediated amplification (RT-LAMP) technology [14]. We identified highly conserved regions of the ZIKV genome and designed RT-LAMP primers for the Zika lineage that is prevalent in the Americas. To enable POC molecular diagnostics, we developed a disposable microfluidic cassette (Fig. 1a) that combines viral nucleic acid capture, concentration, isothermal amplification; and detection. Our disposable, microfluidic cassette contains multiple independent amplification reactors, each equipped with a silica-based nucleic acid isolation membrane at its inlet. The advantage of such a design is to decouple the sample volume from the reaction volume, allowing one to use relatively high sample volumes to achieve high sensitivity. Nucleic acids captured by the isolation membrane directly serve as templates in an RT-LAMP amplification process without a need for an elution step, significantly simplifying flow control.
Our microfluidic cassette can be incubated with battery power or operate electricity-free. To eliminate the need for electricity, we used a simple, thermally insulated portable cup (Fig. 1b) heated by an exothermic reaction for chip-based isothermal amplification [14]. One Mg−Fe alloy pouch, which is usually used as a heater of MRE (meal, ready-to-eat), served as the heat source. Tap water was introduced into the drawer, housing the Mg-Fe pouch, through a port in the cup lid to interact with the Mg−Fe alloy to produce heat. To isolate the amplification reactor’s temperature from variable ambient conditions, we used a phase change material (PCM) to regulate the temperature, removing the need for a thermal control circuit. An aluminium heat sink was used to enhance heat transfer from the PCM to the cassette.
We tested the utility of our POC diagnostic system with raw saliva samples spiked with various concentrations of the ZIKV. Our experiments showed that our electricity-free POC diagnostic system could detect ZIKV in saliva with the sensitivity of 5 plaque forming units (p.f.u.) of ZIKV per sample within 40 min (Fig. 1c). Our POC diagnostic system is comparable to that of the benchtop assay without a need for laboratory facilities, expensive equipment and well-trained personnel [14].
Conclusion
Zika molecular diagnostics can be performed at the point of care with a low-cost, portable system based on a microfluidic cassette that integrates nucleic acid isolation and concentration, isothermal amplification, and detection. To achieve electricity-free isothermal amplification, the cassette is combined with a chemically heated cup that generates heat with an exothermic reaction. The platform can be adapted to various sample types and sizes, and multiplex detection. This flexibility is useful in view of the evolving understanding of Zika pathology and Zika biomarker levels and their persistence in different body fluids and tissues. In the future, we plan to expand system capabilities to enable concurrent detection of multiple vector-borne diseases [15]. Our system is very suitable for resource-poor settings, where funds, centralized laboratory facilities and trained personnel are in short supply, as well as for use in remote clinics and at home.
Acknowledgment
The research reported here was supported, in part by the NIH NIDCR R21DE026700, K25AI099160, R01 CA214072, to the University of Pennsylvania.
References
1. Cao-Lormeau VM, Blake A, Mons S, Lastère S, Roche C, et al. Guillain-Barré Syndrome outbreak associated with Zika virus infection in French Polynesia: a case-control study. Lancet 2016; 387(10027): 1531–1539.
2. Tang H, Hammack C, Ogden SC, Wen Z, Qian X, et al. Zika virus infects human cortical neural progenitors and attenuates their growth. Cell Stem Cell 2016; 18(5): 587–590.
3. Driggers RW, Ho CY, Korhonen EM, Kuivanen S, Jääskeläinen AJ, et al. Zika virus infection with prolonged maternal viremia and fetal brain abnormalities. N Eng J Med 2016; 374: 2142–2151.
4. Kindhauser MK, Allen T, Frank V, Santhana RS, Dye C. Zika: the origin and spread of a mosquito-borne virus. Bull World Health Organ 2016; 94: 675–686C.
5. Lee BY, Alfaro-Murillo JA, Parpia AS, Asti L, Wedlock PT, et al. The potential economic burden of Zika in the continental United States. PLOS Neg Trop Dis 2017; 11(4): e0005531.
6. Current Zika product pipeline. World Health Organization (WHO) 2016. http://www.who.int/csr/research-and-development/zika-rd-pipeline.pdf.
7. Charrel, RN, Leparc-Goffart I, Pas S, de Lamballerie X, Koopmans M, Reusken C. State of knowledge on Zika virus for an adequate laboratory response. Bull World Health 2016; 94: 574–584D.
8. New CDC laboratory test for Zika virus authorized for emergency use by FDA. Centers for Disease Control and Prevention (CDC) 2016. https://www.cdc.gov/media/releases/2016/s0226-laboratory-test-for-zika-virus.html.
9. Zika rapid test. Biocan Diagnostics Inc. http://www.zikatest.com/?page_id=6.
10. Artron Zika test. Artron Laboratories Inc. http://www.artronlab.com/home.html.
11. Gourinat AC, O’Connor O, Calvez E, Goarant C, Dupont-Rouzeyrol M. Detection of Zika virus in urine. Emerg Infect Dis 2015; 21(1): 84.
12. Mansuy JM, Dutertre M, Mengelle C, Fourcade C, Marchou B, et al. Zika virus: high infectious viral load in semen, a new sexually transmitted pathogen. Lancet Infect Dis 2016; 16: 405.
13. Trioplex real-time RT-PCR assay. CDC 2017. https://www.fda.gov/downloads/medicaldevices/safety/emergencysituations/ucm491592.pdf.
14. Song J, Mauk MG, Hackett BA, Cherry S, Bau HH, Liu C. Instrument-free point-of-care molecular detection of Zika virus. Anal Chem 2016; 88: 7289–7294.
15. Song J, Liu C, Mauk MG, Rankin SC, Lok JB, et al. Two-stage isothermal enzymatic amplification for concurrent multiplex molecular detection. Clin Chem 2017; 63(3): 714–722.
The authors
Michael G Mauk PhD, Jinzhao Song PhD, Haim H. Bau PhD, Changchun Liu* PhD
Department of Mechanical Engineering and Applied Mechanics, School of Engineering and Applied Science, University of Pennsylvania, Philadelphia, Pennsylvania 19104, United States
*Corresponding author
E-mail: lchangc@seas.upenn.edu