Preimplantation genetic screening and related issues
Preimplantation genetic screening is a diagnostic approach dedicated to patients undergoing IVF with the proper indications (advanced maternal age, recurrent implantation failure, recurrent pregnancy loss) in order to increase pregnancy rates per transfer via euploid embryo selection. This strategy, and all the associated techniques, are in constant evolution and will shed more light on unexplored aspects of embryology, such as female meiosis or chromosomal mosaicism, creating new criteria for embryo selection.
by Dr D. Cimadomo, Dr A. Capalbo, Dr L. Rienzi and Dr F. M. Ubaldi
Background
Preimplantation genetic diagnosis (PGD) and preimplantation genetic screening (PGS) are two diagnostic approaches increasingly exploited in recent decades within assisted reproduction facilities in the presence of specific indications. PGD is used to identify unaffected embryos in couples at high reproductive risk of a hereditary disease. Usually, these couples conceive naturally and undergo prenatal genetic testing, i.e. villocentesis or amniocentesis; procedures that are invasive and carry a high risk of subsequent miscarriage. The ultimate aim of PGD is, therefore, to prevent the conception of a fetus affected specifically and uniquely by a pathology whose causative mutations have been identified and characterized in the parental genomes before conception. Consequently, PGD depends on a preliminary ad hoc work-up for each couple approaching to an IVF cycle. PGS, instead, is meant to identify only chromosomally normal embryos, thus looking for the presence of chromosomal abnormalities. Since the development of aneuploidies is a de novo event directly linked to maternal age, this diagnostic method is independent from any specific preliminary set-up, thus being identical for each PGS cycle. The indications for this analysis are mainly advanced reproductive maternal age (more than 35 years old; AMA), recurrent implantation failure (more than three failed IVF attempts; RIF) and recurrent pregnancy loss (more than three miscarriages; RPL). From an embryological perspective there is no difference between PGD and PGS. Indeed, strategy and planning of the cycle and biopsy techniques are similar, whereas the genetic technical aspects are significantly different.
Testing for aneuploidy
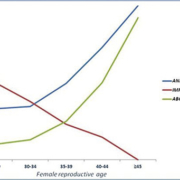
Interestingly, the data collected by the ESHRE PGD consortium IX showed a constant increase in the number of the PGD cycles approached uniquely for euploid embryo selection. In particular, more than 60% of PGD cycles were actually PGS for AMA, RIF or RPL patients, and this percentage is still currently increasing. There is, in fact, a striking impact of aneuploidies on human reproduction. In particular, their incidence in newborns is around 0.3%, mostly represented by trisomies of chromosomes 13, 18 and 21 and sex chromosome aneuploidies. However, tracking backwards through the developmental stages sees this incidence sharply increase, involving other chromosomes and reaching an incidence of up to 60% in preimplantation embryos and 70% in eggs or polar bodies [1]. On the contrary, this incidence in sperm is definitely less severe, as it is never greater than 3–4%. Moreover, a significant number of spontaneous abortions are linked to aneuploidies (more than 60% of products of conception follow chromosomal abnormalities), both increase exponentially with maternal age and fertility rate collapses (Fig. 1) [2].
From a biological perspective, the origin of high trisomy rates found in clinically recognized pregnancies (which sharply increases in patients older than 35 years) resides mainly in maternal meiosis I and II [3]. Recent data obtained through array comparative genomic hybridization (aCGH) on polar bodies (PBs) showed that chromatid errors in female meiosis, such as premature separation of sister chromatids, definitely outnumber impairments involving whole chromosomes as previously thought [4, 5]. Capalbo et al. [5] performed analyses on biopsies at sequential stages of development, in particular the two PBs, a single blastomere at day 3 of embryo development and also a trophectoderm (TE) sample at the blastocyst stage (Fig. 2). This study design allowed the determination of PB analysis accuracy and the impact of male and mitotic errors as well as the evaluation of the occurrence of correction mechanisms throughout preimplantation development. It came to light that 76 out of 78 (97.4%) abnormal meiotic segregations concerned errors involving chromatids rather than whole chromosomes at meiosis I. Furthermore, it unveiled not only a false positive rate in PB biopsy analysis of 20.5%, as just 79.5% (62/78) of meiotic segregation errors identified in PB biopsies were confirmed in blastomeres, but also a false negative rate of 47.6%, as 10 out of 21 embryos showed mitotic or male-derived aneuploidies confirmed at day 3 and at the blastocyst stage of development, which are, obviously, not observable in PBs. This evidence subverts our previous scenario of chromosomal aneuploidy genesis, as well as undermining the reliability of the PB analysis strategy.
Chromosomal mosaicism
From a diagnostic perspective in PGS, post-zygotic mitotic segregation errors are definitely more troubling than meiotic ones, as, whereas the latter involve the same aberrant chromosomal layout in the whole developing embryo, the former entail the phenomenon of chromosomal mosaicism. In the last decade several publications focused on the problem of mosaicism and its influence on PGD/PGS, claiming an incidence fluctuating between 25% [6, 7] and up to more than 70% [8]. Even when these data are analysed with a critical approach, it still emerges that mosaicism is a substantial source of misdiagnosis when the embryo is biopsied at day 3 post-fertilization. This evidence encouraged a shift of the biopsy strategy toward the blastocyst stage and, to this end, different studies were conducted in order to thoroughly describe its cytogenetic constitution and the impact of biopsy itself on embryonic developmental competence. In particular, Capalbo et al. [9] published data outlining the impact of chromosomal mosaicism on a diagnosis at day 5/6 of embryo development as well as the aneuploid cells setting between inner cell mass (ICM) and TE. To this end, a novel method of ICM biopsy was conceived [as described in 9], characterized via KRT18 staining [as described in 10] and its efficiency tested. It led to the absence of TE contamination in 85.7% of the ICM biopsy products, and a low TE contamination rate (only 2% of TE cells) in the rest of them. These data attest the reliability of this biopsy procedure to test the influence of mosaicism at the blastocyst stage. The study design entailed a preliminary aCGH analysis on a TE biopsy during blastocyst-stage PGS clinical cycles, followed by FISH re-analysis of three further fragments of TE and of the ICM from those blastocysts found to be carriers of copy-number chromosomal errors as well as euploid embryos. This revealed that at the blastocyst stage of development, 79.1% of the aneuploidies were constitutional, while 20.9% of them were mosaic. However, only 4% of the blastocysts were found to be mosaic diploid/aneuploid, being at risk of misdiagnosis due to mosaicism when testing at the blastocyst stage. These data strengthen the theory that the impact of mosaicism could be critical at day 3 of embryo development, but it has definitely less influence at the blastocyst stage, thus strongly presenting the latter as the most reliable candidate biopsy stage to perform PGS. Importantly, in the same paper, Capalbo et al. demonstrated that, after excluding low grade mosaicism (<20% of aneuploid cells) and mosaicism confined to one or two TE sections, in 97.1% of cases concordance for all chromosomes re-analysed by FISH between ICM and TE was observed. On a per embryo analysis, instead, complete concordance in TE-based prediction of ICM chromosomal complement was reported (Fig. 3) [9]. Northrop et al. [11] conducted a similar analysis exploiting a single nucleotide polymorphism (SNP) array, which is a comprehensive chromosomal screening technique. This method was found to detect aneuploidy in samples possessing more than 25% aneuploidy, thus when as few as 2 of the 5 cells within a TE biopsy contain the same chromosomal error. Their data showed no preferential aneuploid cell migration to the TE layer, as aneuploidy was observed in 31% of ICM samples (15 out of 48 ICM products) and 32% of TE ones (46 of 144 TE products). Furthermore, a mosaicism rate of 24% was attested, since 12 out of 50 blastocysts screened showed more than a single diagnosis in all of the multiple sections that were re-analysed.
Does the biopsy procedure affect embryo reproductive competence?
One concern about PGS is that biopsy could affect embryo reproductive competence. To investigate this possibility, Scott et al. [12] designed a randomized and paired clinical trial. They selected two of the best quality embryos from the same patient to be transferred and randomized them, one to undergo biopsy, either at day 3 or at day 5 of embryo development, and the other as a control. The biopsy was submitted to SNP array analysis. If only one embryo implanted, buccal DNA obtained from the neonate after delivery was analysed by SNP array to determine whether the implanted embryo was the control one or not. The data collected clearly showed that conducting the biopsy at the cleavage stage affects the clinical outcome, as an absolute reduction in implantation rate of 19.6% with respect to the control was reported. On the contrary, blastocyst biopsy led to a non-significant overall reduction of implantation of 3%; thus an implantation rate equivalent to the control. It is still unclear whether this is due to a smaller proportion of the embryo’s total number of cells being removed, or to the fact that only extra-embryonic cells are involved, or to a higher stress-tolerance of the blastocyst; however, it is still additional important evidence supporting TE biopsy as the ‘gold standard’ for PGS. From a clinical perspective, the same authors also published a randomized controlled trial [13] comparing the clinical outcomes of single euploid blastocyst transfer versus double untested blastocyst transfer. Ongoing pregnancy rates per randomized patient were similar between the two groups (60.7% in the study group vs 65.1% in the control group), whereas a higher multiple pregnancy rate in the control group was recorded (54% vs 0% in the study group). Ultimately then, PGS on TE biopsy associated with a single euploid blastocyst transfer elicits the same clinical outcomes as conventional IVF, but reduces its risks.
Conclusion
In conclusion, PGS is an important diagnostic approach for patients with the proper indications (AMA, RIF or RPL), performed in order to boost implantation rate per transfer. Euploid embryo selection prevents useless and potentially detrimental embryo transfers. Consequently, further advantages of performing PGS are a lower time-to-pregnancy and a higher cost-effectiveness of each single treatment. Moreover, by adopting a biopsy strategy at day 5/6, it is possible to take advantage of a more robust genetic analysis, a high clinical predictive value, the absence of impact of the biopsy on embryo quality, a low influence of mosaicism, as well as a reduced number of embryos to analyse per cycle, as only developmentally competent ones would reach the blastocyst stage. These last aspects will help in reducing costs, thus extending the patients population that can benefit from this technology. Finally, novel comprehensive chromosomal screening techniques, i.e. aCGH, SNP array and quantitative real-time PCR (qPCR), provide us with reliable, sensible and accurate analysis methods, making of PGS also a technically solid approach.
References
1. Nagaoka SI, Hassold TJ, Hunt PA. Human aneuploidy: mechanisms and new insights into an age-old problem. Nat Rev Genet. 2012; 13(7): 493-504.
2. Heffner LJ. Advanced maternal age–how old is too old? N Engl J Med. 2004; 351(19): 1927-1929.
3. Hassold T, Hunt P. To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet. 2001; 2(4): 280-291.
4. Handyside AH, Montag M, Magli MC, Repping S, et al. Multiple meiotic errors caused by predivision of chromatids in women of advanced maternal age undergoing in vitro fertilisation. Eur J Hum Genet. 2012; 20(7): 742-747.
5. Capalbo A, Bono S, Spizzichino L, Biricik A, et al. Sequential comprehensive chromosome analysis on polar bodies, blastomeres and trophoblast: insights into female meiotic errors and chromosomal segregation in the preimplantation window of embryo development. Hum Reprod. 2013; 28(2): 509-518.
6. Voullaire L, Slater H, Williamson R, Wilton L. Chromosome analysis of blastomeres from human embryos by using comparative genomic hybridization. Hum Genet. 2000; 106(2): 210-217.
7. Wells D, Delhanty JD. Comprehensive chromosomal analysis of human preimplantation embryos using whole genome amplification and single cell comparative genomic hybridization. Mol Hum Reprod. 2000; 6(11): 1055-1062.
8. Mertzanidou A, Wilton L, Cheng J, Spits C, et al. Microarray analysis reveals abnormal chromosomal complements in over 70% of 14 normally developing human embryos. Hum Reprod. 2013; 28(1): 256-264.
9. Capalbo A, Wright G, Elliott T, Ubaldi FM, et al. FISH reanalysis of inner cell mass and trophectoderm samples of previously array-CGH screened blastocysts shows high accuracy of diagnosis and no major diagnostic impact of mosaicism at the blastocyst stage. Hum Reprod. 2013; 28(8): 2298-2307.
10. Cauffman G, De Rycke M, Sermon K, Liebaers I, Van de Velde H. Markers that define stemness in ESC are unable to identify the totipotent cells in human preimplantation embryos. Hum Reprod. 2009; 24(1): 63-70.
11. Northrop LE, Treff NR, Levy B, Scott RT Jr. SNP microarray-based 24 chromosome aneuploidy screening demonstrates that cleavage-stage FISH poorly predicts aneuploidy in embryos that develop to morphologically normal blastocysts. Mol Hum Reprod. 2010; 16(8): 590-600.
12. Scott RT Jr, Upham KM, Forman EJ, Zhao T, Treff NR. Cleavage-stage biopsy significantly impairs human embryonic implantation potential while blastocyst biopsy does not: a randomized and paired clinical trial. Fertil Steril. 2013; 100(3): 624-630.
13. Forman EJ, Hong KH, Ferry KM, Tao X, et al. In vitro fertilization with single euploid blastocyst transfer: a randomized controlled trial. Fertil Steril. 2013; 100(1): 100-107.
The authors
Danilo Cimadomo BSc, Antonio Capalbo* PhD, Laura Rienzi MS, Filippo Maria Ubaldi MS
G.EN.E.R.A. Centre for Reproductive
Medicine, Clinica Valle Giulia, Via G. De Notaris 2b, 00197 Rome, Italy
*Corresponding author
E-mail: capalbo@generaroma.it