Simple, fast and highly sensitive colorimetric detection of Zika virus
Background
An outbreak of Zika virus (ZIKV) in Brazil terrorized the whole world and its explosive spread in the Americas caused the World Health Organization (WHO) to declare it a public health emergency of international concern in February 2016 [1]. This is because ZIKV is a suspected major cause of congenital microcephaly, Guillain-Barré syndrome and other neurologic syndromes [2–4]. ZIKV has a genome consisting of a single-stranded, positive-polarity RNA and belongs to the family Flaviviridae and the genus Flavivirus. Aedes mosquitoes, known as a major ZIKV vector, also transmit dengue and chikungunya viruses across tropical and subtropical regions around the world [5]. Moreover, antigenic similarity between ZIKV and dengue virus gives rise to serological cross-reactivity, precluding antibody-based assays from reliably distinguishing between ZIKV and dengue virus infections [6]. Thus, reliable methods for distinguishing ZIKV from dengue and chikungunya viruses are necessary in practical applications.
WHO target product profiles
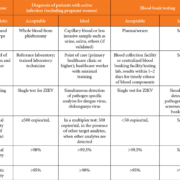
In April 2016, the WHO announced Target Product Profiles (TPPs) for a better diagnostic test for ZIKV infection. The TPPs define the desired characteristics of a ZIKV diagnostic test. The proposed TPPs consist of ‘Detection of active infection with ZIKV’ (Table 1a) and ‘Detection of evidence of prior infection’ (Table 1b). Each characteristic in the tables represents essential properties that the newly developed ZIKV diagnostic test should have at least at an acceptable level. To state the obvious, the criteria of specificity for active infection are more stringent [7].
Previous research on ZIKV diagnostics
Due to serological cross-reactivity between ZIKV and other flaviviruses, most of previous studies on ZIKV diagnosis have dealt with molecular diagnostics instead of immunological assays. Faye and colleagues developed and evaluated a one-step reverse transcription (RT)-PCR assay for ZIKV detection. The limit of detection of the assay was found to be 7.7 plaque-forming units (p.f.u.) per reaction in human serum and in the L-15 medium [8]. A quantitative real-time RT-PCR assay for ZIKV was also developed by the same research group. Analytical sensitivity of the assay was estimated at 3.2×102 RNA copies/μL [9]. However, a conventional PCR assay requires a bulky and expensive thermal cycler, prolonged reaction time, and trained technicians; these resources are not available in many low- and middle-income countries. Moreover, the RT-PCR reaction is vulnerable to inhibitors (blood, plasma and urine), thus requiring painstaking and cumbersome RNA extraction steps.
Recent research on ZIKV diagnostics
To overcome such limitations of RT-PCR, a variety of isothermal nucleic acid amplification techniques have recently been developed. Among them, reverse transcription loop-mediated isothermal amplification (RT-LAMP) is a rapid, robust, and highly sensitive isothermal RNA amplification method that uses four to six primers to amplify specific RNA sequences at 60–65°C even in the presence of inhibitors such as blood, plasma, or urine. RT-LAMP is much faster than conventional PCR, and the reaction can even proceed in an oven, water bath or with heating packs [10, 11]. Despite these advantages, the RT-LAMP assays still rely on a conventional bulky amplicon analyser such as a gel electrophoresis apparatus or a fluorescence laser-induced detector for monitoring the LAMP amplicons; this situation precludes the use of RT-LAMP in point-of-care diagnosis.
Our approaches to simple and highly sensitive diagnosis of ZIKV
To eliminate the dependence on a conventional amplicon analyser while retaining the aforementioned advantages of RT-LAMP, we selected the lateral flow assay (LFA) format for RT-LAMP amplicon analysis. The LFA, a driving principle behind pregnancy test strips, is also widely known as a superior diagnostic tool for nucleic acids owing to its high sensitivity, simplicity, selectivity and easy interpretation of results. Moreover, the Bst 3.0 polymerase used in this study for RT-LAMP retains both improved isothermal amplification performance and strong reverse transcription activity, allowing us to avoid addition of exogenous reverse transcriptase and the inhibition of reverse transcription by biological substances. By utilizing the advantages of Bst 3.0 polymerase and combining the RT-LAMP assay with the LFA, we demonstrated simple and highly sensitive detection of ZIKV RNA in human whole blood by merely observing a colorimetric signal within 35 min.
The RT-LAMP reaction and modification of amplicons in our study
As mentioned above, RT-LAMP has excellent tolerance to many inhibitors so that isothermal amplification of ZIKV RNA is possible even when human whole blood is directly used as a sample. We extracted ZIKV RNA and added it into human whole blood to mimic ZIKV-containing blood samples. Then, the spiked human whole blood was serially diluted with blood to set up a concentration range from 106 copies of RNA to a single copy per 2 μL and directly used these dilutions as samples without additional RNA purification steps. To colorimetrically detect the result of the LFA, a special modification is needed: labelling of the amplicon with digoxigenin and biotin. Among our own designed ZIKV-specific primers, two loop primers were tagged with digoxigenin at the 5´end; this approach will allow digoxigenin to label the amplicon when loop primers amplify the ZIKV RNA by the LAMP method. Labelling of the amplicon with biotin is made possible by adding biotin-labelled dUTP (Biotin-dUTP) to the mix of deoxynucleotides (dNTPs) at a certain ratio. When ZIKV RNA is amplified and this reaction consumes dNTPs, Biotin-dUTP will substitute thymine at the adenine sites of the complementary strand, resulting in labelling of the amplicon with biotin.
RT-LAMP was carried out in a 25 μL reaction mixture containing 1× Isothermal Amplification Buffer II [20 mM Tris-HCl, 10 mM (NH4)2SO2, 150 mM KCl, 2 mM MgSO4, and 0.1% Tween 20], additional 2 mM MgSO4, a dNTP mix supplemented with biotin-dUTP (2.2 mM dGTP, dATP, dCTP, 1.375 mM dTTP, and 0.0825 mM biotin-dUTP), a target-specific primer mixture (0.8 μM forward and reverse inner primers, 0.4 μM digoxigenin-labelled loop primers, and 0.2 μM forward and reverse outer primers), 8 U of Bst 3.0 DNA polymerase, and 2 μL of human whole blood spiked with ZIKV RNA ranging from 106 copies to a single copy per 2 μL. The RT-LAMP reaction mixture was incubated for 30 min.
Design and operation of the LFA
Figure 1(a) and 1(b) shows the detailed set-up and operating procedures of the LFA in our study. First, 1 μL of digoxigenin- and biotin-labelled RT-LAMP products was loaded onto the conjugate pad, so that the biotin-labelled RT-LAMP products formed a complex with gold nanoparticles (AuNPs) via streptavidin-biotin interactions. Next, 45 μL of diluent buffer was placed on the buffer loading pad, and then capillary flow transferred AuNPs from the conjugate pad to the test and control line. The AuNP–RT-LAMP complexes were immobilized at the test line by the interaction between digoxigenin and anti-digoxigenin whereas the AuNPs that did not form complexes were captured by biotin. Complexed and uncomplexed AuNPs are indicated by violet bands at the test line and control line, respectively. The colorimetric signal was easily visible with the naked eye within 5 min.
Discussion
Analysis of the limit of detection in human whole blood samples
We evaluated the limit of detection of the LFA to determine whether our method is indeed highly sensitive. Two microliters of human whole blood was directly used as a sample without any purification steps. Figure 1(c) shows the ZIKV RNA detection results for the LFA. The signal intensities on the test line gradually declined as the concentration of ZIKV RNA decreased. Notably, the presence of even a single copy of ZIKV RNA could be detected within 35 min by the LFA. These results imply that our method has a great potential for diagnosis of ZIKV infections.
References
1. Lessler J, Chaisson LH, Kucirka LM, Bi Q, Grantz K, Salje H, et al. Assessing the global threat from Zika virus. Science 2016; 353: aaf8160.
2. Schuler-Faccini L, Ribeiro E, Feitosa I, Horovitz D, Cavalcanti D, et al. Possible Association Between Zika Virus Infection and Microcephaly Brazil, 2015. MMWR Morb Mortal Wkly Rep 2016; 65: 59–62.
3. Cao-Lormeau V-M, Blake A, Mons S, Lastère S, Roche C, et al. Guillain-Barré Syndrome outbreak associated with Zika virus infection in French Polynesia: a case-control study. Lancet 2016; 387: 1531–1539.
4. WHO statement on the first meeting of the International Health Regulations (2005) (IHR 2005) Emergency Committee on Zika virus and observed increase in neurological disorders and neonatal malformations. http://www.who.int/mediacentre/news/ statements/2016/1st-emergency-committee-zika/en/ (accessed May 1).
5. Surveillance and Control of Aedes aegypti and Aedes albopictus in the United States. http://www.cdc.gov/chikungunya/resources/ vector-control.html (accessed May 1).
6. Dejnirattisai W, Supasa P, Wongwiwat W, Rouvinski A, Barba-spaeth G, et al. Nat Immunol 2016; doi:10.1038/ni.3515.
7. Target product profiles for better diagnostic tests for Zika virus infection. http://www.who.int/csr/research-and-development/zika-tpp.pdf.
8. Faye O, Faye O, Dupressoir A, Weidmann M, Ndiaye M, Alpha Sall A. One-step RT-PCR for detection of Zika virus. J Clin Virol 2008; 43: 96–101.
9. Faye O, Faye O, Diallo D, Diallo M, Weidmann M, Sall AA. Quantitative real-time PCR detection of Zika virus and evaluation with field-caught mosquitoes. Virol J 2013; 10: 311.
10. Safavieh M, Kanakasabapathy MK, Tarlan F, Ahmed MU, Zourob M, et al. Emerging loop-mediated isothermal amplification-based microchip and microdevice technologies for nucleic acid detection. ACS Biomater Sci Eng 2016; 2: 278–294.
11. Nyan D-C, Ulitzky LE, Cehan N, Williamson P, Winkelman V, et al. Rapid detection of hepatitis B virus in blood plasma by a specific and sensitive loop-mediated isothermal amplification assay. Clin Infect Dis 2014; 59: 16–23.
The authors
Dohwan Lee MS, Yong Kyoung Yoo PhD, and Jeong Hoon Lee* PhD
Department of Electrical Engineering, Kwangwoon University, Nowon, Seoul 01897, Republic of Korea.
*Corresponding author
E-mail: jhlee@kw.ac.kr