The challenge of early diagnosis of rheumatoid arthritis patients: looking for better biomarkers
By F..L. Ochoa-González, J..C. Fernández-Ruiz, M..F. Romo-García and Dr J..E. Castañeda-Delgado
Rheumatoid arthritis shows a prevalence of 0.5–1.6 % globally. The identification of biomarkers for early treatment response could aid in the fine tuning of therapy and therefore contribute to increased treatment efficacy and the timely use of biologicals when no response from disease-modifying anti-rheumatic drugs is observed. Our group has identified several biomarkers for early diagnosis and treatment response.
Introduction
Rheumatoid arthritis (RA) is a chronic disease of autoimmune etiology characterized by persistent inflammation of the synovial membrane, which lines the inner surface of capsules of synovial joints. The worldwide estimated prevalence is about 1 % of the adult population. It is more frequent in women with a ratio of 3:1. The cause of RA is unknown; however, genetic and environ-mental factors contribute to RA. Several genes have been associated with an increased risk of developing RA: mainly certain HLA class II antigens associated with the shared epitope that is responsible for antigen presentation to lymphocytes. Smoking and some causative microorganisms of oral diseases such as periodontitis and gingivitis (Porphyromonas gingivalis and Aggregati-bacter Actimomyctemcomitans) have also been associated with RA [1]. The relation-ship of genetic traits/environment and the link to inflammation and autoimmunity is being explored. In this regard post-translational modifications of proteins such as citrullination of arginine by peptidyl arginine deiminase (PAD; some of it mediated by PAD-like enzymes coming from oral pathogens) or carbamylation of lysine (mediated by cyanate from cigar smoke) contribute to breaking immunological tole-rance by creating neoepitopes of autologous proteins resulting in generation of auto-antibodies against modified peptides. An examples include anti-citrullinated protein antibodies (ACPAs), antibodies to the Fc part of IgG [rheumatoid factor (RF)], or autoantigens that cross-react with bacterial or viral antigens [2]. These autoantibodies contribute to the increased inflammatory response observed in RA patients.
The clinical manifestations of RA are mainly associated with symmetric inflammation of small and large joints, accompanied by morning stiffness. Patients with RA usually present multiple comorbidities as a result of chronic inflammation, the main ones being cardiovascular disease or pulmonary mani-festations. RA greatly affects the patient’s quality of life, as it interferes with physical function. In long-term disease without treatment, the accumulation of joint damage is irreversible and leads to disability at an early age without the possibility of recovering normal function. Therefore, it is of great importance to establish an early diagnosis; it has already been shown that beginning treatment prevents the progression of joint damage in up to 90 % of patients in the early stages of the disease.
There is no cure for RA, which is why the goal of treatment is to reach remission, defined as no disease activity and low disease activity with low risk to progression. Therefore, therapeutic approaches are based on drugs that interfere with signs and symptoms of RA, such as disease-modifying antirheumatic drugs (DMARDs). DMARDs are categorized into conventional synthetic (csDMARDs), targeted synthetic (tsDMARDs) and biologic (bDMARDs). The csDMARDs include sulfasalazine, leflunomide, hydroxychloroquine and methotrexate. The joint working group of the American College of Rheumatology and the European League Against Rheumatism (ACR-EULAR) recommends treating all new cases of RA as soon as possible using methotrexate combined with short-term glucocorticoids. It has been reported that a proportion of patients treated with either DMARDs or bDMARDs do not reach treatment target (reduction of DAS28 and disease activity). The presence of autoantibodies, joint damage and high disease activity are associated with rapid disease progression that can be slowed by adding bDMARDs [3]; therefore, better prognostic markers for treatment response are also needed.
According to current ACR-EULAR 2010 classification criteria (not diagnostic criteria), RA patients have joint pain and synovial inflammation, morning stiffness in the joints with duration of at least 30 minutes. There are several serological determinations that aid in the classification of these patients, such as the use of cyclic citrullinated peptides (CCPs) to detect ACPAs, as well as RF and erythrocyte sedimentation rate [4]. Nevertheless, the diagnostic tests, such as the CCP test, that have a high specificity (range: 90–96 %) have several caveats: (1) low sensitivity (range 67–83 %); and (2) when negative, RA cannot be discarded because of the possibility that the patient has seronegative RA [5]. This is also the case for RF, which shows a similar sensitivity but a lower specificity. Therefore, there is a need for diagnostic tools that could help to further clarify the diagnosis of RA. As stated above, early diagnosis and treatment remains one of the crucial points for the management of RA and thus prevention of loss of physical function; however, the auxiliary diagnostic tools remain insufficient to distinguish RA from other rheumatic diseases. It is for these reasons that our group has focused on the search for early biomarkers of disease and of treatment response.
MicroRNAs as diagnostic biomarkers
MicroRNAs (miRNAs) are small RNA molecules (approximately 21 bp long) that modulate transcription and translation. Without miRNAs all the genes that are transcribed into mRNA (messenger RNA) would be translated to proteins, but miRNAs regulate which mRNA will or will not be translated into proteins. One single miRNA can control the production of many proteins and therefore small changes in the abundance of an miRNA could result in bigger changes at the protein level. This is these molecules are very important and are indicators of what is happening inside the body before these changes can be observed as clinical symptoms.
Recently, using microarray technology, our group detected changes in the expression profile of several hundred mRNAs in patients with early RA whose symptoms at that moment were barely classifiable by a rheumatologist [6]. What was behind that drastic change? As miRNAs are the main regulators of mRNA, we hypothesized that miRNAs could be responsible. Therefore, we analysed this miRNA profile in patients with early RA (using the same technology) and we identified 97 miRNAs that were over-expressed in early RA [7]. It seemed (but more experiments are needed to confirm this) that only 97 miRNAs are responsible for regulating around 2000 mRNA and all this in the early phases of the disease and the start of arthritis, when symptoms can’t be clearly classified by the clinician. This discovery encouraged us to explore whether any of these miRNAs can serve as a biomarker for the detection of early RA. Thus we performed an analysis named receiver operator characteristic curve (ROC curve). This kind of graph shows how many patients a biomarker (in this case a miRNA) can classify correctly and how many incorrectly. From this analysis we found that miRNA mir-361-5p had a specificity of 82.61 and sensitivity of 81.25. The value for sensitivity is bigger than the one of CCP (as mentioned previously) meaning that probably mir-361-5p can help to correctly identify those with the disease (true positives). However, a wider study is needed to confirm such observations and take it to the clinical lab.
The interest in miRNAs as biomarkers is increasing not only because they are master regulators as we mentioned before, but also because they have much greater stability in several fluids types, such as saliva or serum [8], compared to mRNA and this facilitates detection. Currently, the detection of miRNAs is performed by PCR or massive sequencing, which are technologies that need special equipment and infrastructure. However, recently, research groups have started to work on novel ideas for miRNA detection, such as small electronic devices to quantify miRNAs from a drop of serum [9].
Circulating miRNAs as potential biomarkers of treatment response
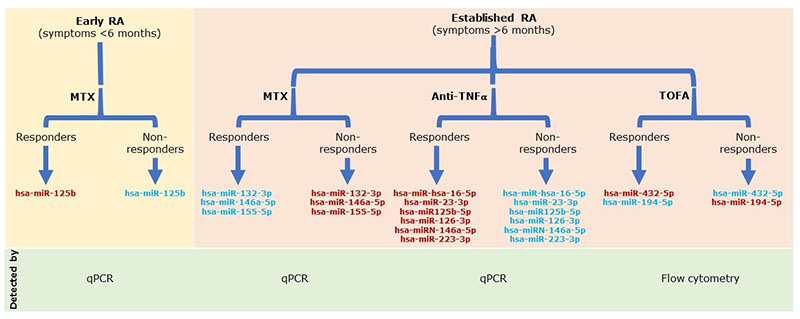
miRNAs have been proposed by different authors as possible biomarkers of response to csDMARDs, bDMARDs and tsDMARDs in RA. The abnormal expression pattern of miRNAs reflects the underlying patho-physiological processes owing to its direct relationship with the inflammatory processes. Some miRNAs have been described as markers of response to anti-tumour necrosis factor alpha (TNFα) treatment in patients with RA, which is the case with hsa-miR-hsa-16-5p, hsa-miR-23-3p, hsa-miR125b-5p, hsa-miR-126-3p, hsa-miRN-146a-5p and hsa-miR-223-3p that are upregulated in patients who respond after therapy and show a reduction in inflammatory parameters [TNFα, interleukin-6 (IL-6), IL-17, RF and C-reactive protein (CRP)] [10, 11]. Even methotrexate treatment seemed to have an effect on the expression of hsa-miR-132-3p, hsa-miR-146a-5p and hsa-miR-155-5p, where good responders have a downregulation of these miRNAs [12] (Fig. 1).
Our research team evaluated the expression of serum miRNAs by flow cytometry (Firefly™ technology) in a cohort of patients receiving treatment with tofacitinib (TOFA, a tsDMARD) for 5 years as part of an open-label study. Treatment with TOFA does not affect miRNA expression directly; however, in patients experiencing RA flare-up we identified changes in two miRNAs: hsa-miR-432-5p was downregulated and hsa-miR-194-5p was upregulated. Our findings suggest that these miRNAs could be used as a biomarkers for relapse. By monitoring them, relapse could be predicted and prevented by allowing a return to a treatment scheme before the patient’s symptoms worsen. How-ever, more research is required, as it is the first time that these miRNAs have been found to be involved in the inflammatory response of patients with RA [13]. If these findings can be confirmed, these miRNAs could be useful biomarkers for prediction of therapy effectiveness as well as therapy monitoring and could, therefore, be a useful support tool for generating personalized treatment regimens for RA patients.
Perspectives
The miRNA biomarkers reported above await further validation in the clinical setting. The design of clinical studies for such validation should account for and analyse possible reallife situations, such as the similarity of symptoms of RA patients with other rheumatic diseases and confounding variables.
References
1. McInnes IB, Schett G. The pathogenesis of rheumatoid arthritis. N Engl J Med 2011; 365: 2205–2219.
2. Cheng Z, Meade J, Mankia K, Emery P, Devine DA. Periodontal disease and periodontal bacteria as triggers for rheumatoid arthritis. Best Pract Res Clin Rheumatol 2017; 31: 19–30.
3. Calabrese LH, Calabrese C, Kirchner E. The 2015 American College of Rheumatology Guideline for the treatment of rheumatoid arthritis should include new standards for hepatitis B screening: comment on the article by Singh et al. Arthritis Care Res 2016; 68: 723–724.
4. Kay J, Upchurch KS. ACR/EULAR 2010 rheumatoid arthritis classification criteria. Rheumatology 2012; 51(Suppl 6): vi5–9.
5. Mathsson Alm L, Fountain DL, Cadwell KK, Madrigal AM, Gallo G, Poorafshar M. The performance of anti-cyclic citrullinated peptide assays in diagnosing rheumatoid arthritis: a systematic review and meta-analysis. Clin Exp Rheumatol 2018; 36: 144–152.
6. Macías-Segura N, Castañeda-Delgado JE, Bastian Y, Santiago-Algarra D, Castillo-Ortiz JD, Alemán-Navarro AL, Jaime-Sánchez E, Gomez-Moreno M, Saucedo-Toral CA, et al. Transcriptional signature associated with early rheumatoid arthritis and healthy individuals at high risk to develop the disease. PLoS One 2018; 13: e0194205.
7. Romo-García MF, Bastian Y, Zapata-Zuñiga M, Macías-Segura N, Castillo-Ortiz JD, Lara-Ramírez EE, Fernández-Ruiz JC, Berlanga-Taylor AJ, González-Amaro R, et al. Identification of putative miRNA biomarkers in early rheumatoid arthritis by genome-wide microarray profiling: A pilot study. Gene 2019; 720: 144081.
8. Huang W. MicroRNAs: biomarkers, diagnostics, and therapeutics. Methods Mol Biol 2017; 1617: 57–67.
9. Labib M, Khan N, Ghobadloo SM, Cheng J, Pezacki JP, Berezovski MV. Three-mode electrochemical sensing of ultralow microRNA levels. J Am Chem Soc 2013; 135: 3027–3038.
10. Castro-Villegas C, Pérez-Sánchez C, Escudero A, Filipescu I, Verdu M, Ruiz-Limón P, Aguirre MA, Jiménez-Gomez Y, Font P, Rodriguez-Ariza A, et al. Circulating miRNAs as potential biomarkers of therapy effectiveness
in rheumatoid arthritis patients treated with anti-TNFα. Arthritis Res Ther 2015; 17: 49.
11. Filková M, Aradi B, Senolt L, Ospelt C, Vettori S, Mann H, Filer A, Raza K, Buckley CD, et al. Association of circulating miR-223 and miR-16 with disease activity in patients with early rheumatoid arthritis. Ann Rheum Dis 2014; 73(10): 1898–1904.
12. Singh A, Patro PS, Aggarwal A. MicroRNA-132, miR-146a, and miR-155 as potential biomarkers of methotrexate response in patients with rheumatoid arthritis. Clin Rheumatol 2019; 38: 877–884.
13. Fernández-Ruiz JC, Ramos-Remus C, Sánchez-Corona J, Castillo-Ortiz JD, Castañeda-Sánchez JJ, Bastian Y, Romo-García MF, Ochoa-González F, Monsivais-Urenda AE, et al. Analysis of miRNA expression in patients with rheumatoid arthritis during remission and relapse after a 5-year trial of tofacitinib treatment. Int Immunopharmacol 2018; 63: 35–42.
The authors
Fatima de Lourdes Ochoa-González1,2 MSc, Julio Cesar Fernández-Ruiz2,3 MSc, Maria Fernanda Romo García2,3 MSc, Julio Enrique Castañeda-Delgado*4 PhD
1 Unidad Académica de Biología, Universidad Autónoma de Zacatecas, Zacatecas, México
2 Unidad de Investigación Biomédica de Zacatecas, Instituto Mexicano del Seguro Social, Zacatecas, México
3 Centro de Investigación en Ciencias de la Salud y Biomedicina, Universidad Autónoma de San Luís Potosí, San Luís Potosí, México
4 Cátedras-CONACYT, Unidad de Investigación Biomédica de Zacatecas, Instituto Mexicano del Seguro Social, Zacatecas, México
*Corresponding author
E-mail: julioenrique_castaneda@yahoo.com.mx
Figure 1. Circulating microRNAs after treatment for rheumatoid arthritis (RA). Flow diagram showing the expression of some microRNAs after methotrexate (MTX), anti-TNFα and tofacitinib (TOFA) treatment. Red, upregulated microRNAs; blue, downregulated microRNAs; qPCR, quantitative PCR.
…