Tissue biomarkers of breast cancer: implications for prognosis
Better tissue biomarkers are needed to improve diagnosis and prognosis, guide molecularly targeted therapy, and monitor activity and therapeutic response across many cancers. Proteomics methods, based on mass spectrometry, hold great promise for the discovery of novel biomarkers that might form the foundation of a new clinical test. This review will focus on potential tissue biomarkers with utility for prognosis in breast cancer.
By Dr Liping Chung
Tissue biomarkers in breast cancer
Breast cancer is the leading cause of mortality among women worldwide. It is a complex and heterogeneous disease and includes several subtypes, which have different prognoses and responses to therapy. Recent molecular characterization of some breast cancer subtypes has led to the development of personalized options for treatment targeting [1].
One of the major advantages of biomarker research for individuals with cancer is the availability of tumour tissue for analysis and the possibility that potential tissue biomarkers can be detected in histological samples. In conjunction with tumour grading and measurement of lymphovascular invasion, several tissue biomarkers are now used with prognostic significance in daily practice including estrogen receptor (ER), progesterone receptor (PR), the type 2 epidermal growth factor receptor (HER2 or erbB-2), and Ki67 [1, 2].
The identification of protein biomarkers in easily accessible biological fluids has potential for the development of minimally invasive procedures for early diagnostics, but the analysis of body fluids such as plasma, serum and urine is complicated by their wide dynamic range of protein expression, the variation in their composition and their sensitivity to sample handling. Many serum biomarkers are not very specific or sensitive [1]. Analysis of tissue homogenates using the well-established and extremely powerful conventional techniques of differential proteomics has the advantage of covering the lower range of protein expression in such samples than in biological fluids [3].
Prognosis and response prediction
Different from diagnostic markers that detect the potential for developing a malignancy or test for the presence of a malignancy, biological markers that predict prognosis once a cancer has occurred are of great importance because they may influence major therapeutic recommendations. For breast cancer, these markers have become part of contemporary clinical practice. Among established tissue marker proteins in breast cancer, ER and HER2 are not diagnostic but have the greatest predictive utility [2]. It is generally accepted that estrogen receptor-positive (ER+) and ER-negative (ER−) breast cancers represent different disease entities. ER- tumours tend to be of high grade, have more frequent p53 mutations, and have worse prognosis compared with ER+ disease. Both ER+ and ER- tumours can be either HER2 positive or negative. Low-grade tumours are typically ER positive, and almost always HER2 non-amplified. The approximately 15% of patients with breast cancer who have HER2 overexpressing and amplified tumours are typically treated with a combination of trastuzumab, a monoclonal antibody targeting HER2, and adjuvant chemotherapy [4]. HER2 amplification and overexpression are generally associated with a poor prognosis. The prognostic significance of HER2 overexpression in tumour tissue has been evaluated in several clinical trials, suggesting that HER2 positivity is correlated with worse prognosis in untreated breast cancer patients, including node-negative populations [5].
The search for breast tissue biomarkers by mass spectrometry-based proteomics
Proteomic approaches, particularly those involving mass spectrometry (MS), have been widely used in breast cancer biomarker discovery, although to date no new markers based on proteomic discovery have been adopted for use in clinical practice. Using laser capture microdissection (LCM) for tissue samples, an extensive tissue study was performed by MALDI-MS (matrix-assisted laser desorption/ionization mass spectrometry) analysis on an average of 2000 cells from 122 invasive mammary carcinomas and 167 samples of normal breast epithelium [6]. Among clusters of protein/peptide peaks that were used to discriminate cancer from normal tissue with high sensitivity and specificity were ubiquitin, S100A6 (calcyclin) and S100A8 (calgranulin A). To confirm cDNA expression profiling of breast tissues, Brozkova et al. also analysed whole tissue lysates rather than serum of 105 breast carcinomas on IMAC30 protein chips by SELDI-TOF MS (surface-enhanced laser desorption/ionization, time-of-flight mass spectrometry) [7]. They compared this analysis to cDNA expression profiling of the same tumours and found similar clustering, providing supporting evidence for the effectiveness of this technique in identifying and classifying tumours.
Most clinical tissue samples are conserved as formalin-fixed paraffin-embedded (FFPE) samples. In particular, cancer tissues contain several different cell types at various developmental stages. It was generally believed that proteins in FFPE tissues were altered and inaccessible for analysis by mass spectrometry until recent developments have shown it is possible to access the protein in imaging mass spectrometry (IMS) experiments following antigen retrieval [8]. The direct analysis of cancer tissues by IMS preserves the spatial proteomic information. Consequently, it is holds great promise for the discovery of highly specific biomarkers. A recent study demonstrated the potential of MALDI-imaging MS for HER2 status of clinical parameters in cases of breast cancer based on protein patterns. This potentially allows the selection of patients likely to respond to trastuzumab treatment. Comparing the HER2-positive (HER2+) vs HER2-negative (HER2−) breast cancer protein profiles, the authors found a specific proteomic signature of seven species, able to accurately classify the HER2 status with a sensitivity of 83%, a specificity of 92% and an overall accuracy of 89% [9].
Protein biomarkers and conventional pathologic features
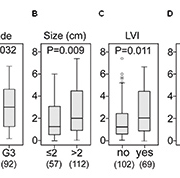
In a very recent study, using protein extracts of breast tissues (n=171), we have used SELDI-TOF MS to discover two proteins that, in combination, show high discrimination between breast cancer and healthy breast tissue samples [10]. These putative breast cancer biomarkers were verified on an independent sample set, and identified as ubiquitin and a novel truncated form of the S100 protein family member, S100P. Interestingly, the combined panel of two protein markers was significantly associated with tumour histologic grade, size, and lymphovascular invasion (LVI), and also with ER-positive (ER+) and PR-positive (PR+) status and HER2 overexpression. In particular, as shown in Figure 1, significant positive associations were seen between a previously unreported short form of S100P (9.2kDa) and tumour size, high grade, LVI and lymph node involvement (LN), and also associated with hormone receptor positive status and HER2 overexpression (unpublished data). These results implicate that a protein biomarker panel may indicate a HER2-enriched breast cancer subtype with poor prognosis, and that measurement of S100P may be valuable both in the classification of breast cancer and as a possible target for treatment. Furthermore, in another very recent study, the prognostic value of S100P was also tested for FFPE tissue obtained from 85 breast cancer patients with a median follow up of 17 years. High immunocytochemical staining of breast tumour sections for S100P has been associated with poor long-term patient survival [11].
Conclusion and future prospects
In this era of using new high-throughput methods, many new protein biomarkers have been reported for both prognostic and predictive purposes. However, none of these have been widely accepted in routine clinical practice, possibly due to a lack of sufficient validation to meet the criteria of the American Society of Clinical Oncology’s tumour marker utility grading system and guideline recommendations [1]. Identification of novel markers based on gene expression and proteomic profiling has led to more definitive insights into tumour biology. The accurate evaluation of the status of clinical parameters in cases of breast cancer is of primary importance for prognostic value and therapeutic decision. Different methodologies successfully used for breast cancer prognostic information and therapy outcome prediction may suggest that the future diagnostics and consequent individualization of therapy will become much more wide-ranging.
References
1. Harris L, Fritsche H, Mennel R, Norton L, Ravdin P, Taube S, Somerfield MR, Hayes DF, Bast RC, Jr. American Society of Clinical Oncology 2007 update of recommendations for the use of tumour markers in breast cancer. J Clin Oncol 2007; 25(33): 5287–5312.
2. Chung L, Baxter RC. Breast cancer biomarkers: proteomic discovery and translation to clinically relevant assays. Expert Rev Proteomics 2012; 9(6): 599–614.
3. Danova M, Delfanti S, Manzoni M, Mariucci S. Tissue and soluble biomarkers in breast cancer and their applications: ready to use? Journal of the National Cancer Institute Monographs 2011; 2011(43): 75–78.
4. Cheang MC, Chia SK, Voduc D, Gao D, Leung S, Snider J, Watson M, Davies S, Bernard PS, Parker JS, et al. Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. Journal of the National Cancer Institute 2009; 101(10): 736–750.
5. Andrulis IL, Bull SB, Blackstein ME, Sutherland D, Mak C, Sidlofsky S, Pritzker KP, Hartwick RW, Hanna W, Lickley L, et al. neu/erbB-2 amplification identifies a poor-prognosis group of women with node-negative breast cancer. Toronto Breast Cancer Study Group. J Clin Oncol 1998; 16(4): 1340–1349.
6. Sanders ME, Dias EC, Xu BJ, Mobley JA, Billheimer D, Roder H, Grigorieva J, Dowsett M, Arteaga CL, Caprioli RM. Differentiating proteomic biomarkers in breast cancer by laser capture microdissection and MALDI MS. J Proteome Res 2008; 7(4): 1500–1507.
7. Brozkova K, Budinska E, Bouchal P, Hernychova L, Knoflickova D, Valik D, Vyzula R, Vojtesek B, Nenutil R. Surface-enhanced laser desorption/ionization time-of-flight proteomic profiling of breast carcinomas identifies clinicopathologically relevant groups of patients similar to previously defined clusters from cDNA expression. Breast Cancer Res 2008; 10(3): R48.
8. Gustafsson JO, Oehler MK, McColl SR, Hoffmann P. Citric acid antigen retrieval (CAAR) for tryptic peptide imaging directly on archived formalin-fixed paraffin-embedded tissue. J Proteome Res 2010; 9(9): 4315–4328.
9. Rauser S, Marquardt C, Balluff B, Deininger SO, Albers C, Belau E, Hartmer R, Suckau D, Specht K, Ebert MP, et al. Classification of HER2 receptor status in breast cancer tissues by MALDI imaging mass spectrometry. J Proteome Res 2010; 9(4): 1854–1863.
10. Chung L, Shibli S, Moore K, Elder EE, Boyle FM, Marsh DJ, Baxter RC. Tissue biomarkers of breast cancer and their association with conventional pathologic features. Br J Cancer 2013; 108(2): 351–360.
11. Maciejczyk A, Lacko A, Ekiert M, Jagoda E, Wysocka T, Matkowski R, Halon A, Gyorffy B, Lage H, Surowiak P. Elevated nuclear S100P expression is associated with poor survival in early breast cancer patients. Histol Histopathol 2013; 28(4): 513–524.
The author
Liping Chung PhD
Kolling Institute of Medical Research,
University of Sydney, Royal North Shore Hospital, NSW 2065, Australia
E-mail: liping.chung@sydney.edu.au