Indirect immunofluorescence (IIF) is an indispensable method for autoantibody diagnostics, providing high sensitivity and specificity together with a broad antigenic spectrum. However, the microscopic evaluation of the fluorescence patterns is both time-consuming and challenging for laboratory staff, and is, moreover, based on subjective interpretation. Laboratories are increasingly turning to automated systems to facilitate and standardize the IIF readout and interpretation. In recent years various automation systems have been developed, which provide automated digital acquisition of IIF images, discrimination of positive and negative samples, as well as pattern classification for key applications. This article focuses on the EUROPattern system, which provides computer-aided immunofluorescence microscopy for anti-nuclear antibodies (ANA), anti-neutrophil granulocyte cytoplasm antibodies (ANCA), antibodies against double-stranded DNA (anti-dsDNA) on Crithidia luciliae, monospecific antigen microdots (EUROPLUS) and transfected cell-based assays e.g. for anti-neuronal antibodies. The accuracy of automated evaluation compared to visual assessment has been investigated in various published studies.
by Dr Jacqueline Gosink
ANA
ANA represent a key diagnostic criterion for many autoimmune diseases, including systemic lupus erythematosus (SLE), mixed connective tissue disease, Sjögren’s syndrome, systemic sclerosis, polymyositis, dermatomyositis and primary biliary cirrhosis. The gold standard for ANA determination is IIF on human epithelial (HEp-2) cells. This substrate provides the complete antigen spectrum and allows investigation of over 100 different autoantibodies. Observation of the fluorescence pattern enables classification of the antibody or antibodies present in the patient sample. Positive results are confirmed by monospecific tests such as ELISA, immunoblot or IIF microdot assays.
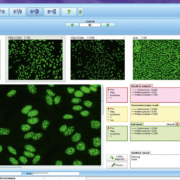
The automated evaluation of HEp-2 cells includes reliable discrimination of positive and negative ANA results, as well as classification of all ANA patterns [1] (Figure 1), encompassing homogeneous, speckled, nuclear dots, nucleolar, centromeres, nuclear envelope and cytoplasmic. The ANA patterns identified by EUROPattern correspond to the competent level reporting defined by the International Consensus on ANA Patterns (ICAP; www.anapatterns.org). Mixed patterns, which occur when more than one antibody is present, are also recognized and reported as such. The pattern is assigned by analysing its features and comparing it to a reference database of over 5000 images, corresponding to 115,000 cells. Unspecific signals originating from outside of the cells are identified by means of a DNA counterstain and subsequently rejected. The evaluation also includes titre designations with confidence values for the detected antibodies. Results from the HEp-2 screening can be monospecifically confirmed using microdot substrates of purified antigens, which are incubated and evaluated in parallel.
To assess the diagnostic accuracy, the automated evaluation was compared to conventional visual interpretation by experts in the field using 351 patient sera [2]. The concordance for positive/negative discrimination was 99%, with an analytical sensitivity of 100% and a specificity of 98%. In 60% of samples, the pattern, including variable mixed patterns, was recognized completely by the software. In 94% of samples, the main pattern was correctly designated. A further study showed 79% correct pattern assignment.
Anti-dsDNA antibodies
Anti-dsDNA antibodies are a hallmark of SLE and represent an important criterion for diagnosis. Their prevalence in SLE ranges from 30% to 98% in different studies, depending among other things on the test method used. Like the gold standard Farr assay, IIF using Crithidia luciliae as the substrate (CLIFT) is considered to have a very high disease specificity. The method takes advantage of the kinetoplast of C. luciliae, which is rich in DNA but contains hardly any other antigens, thus enabling highly selective detection of anti-dsDNA antibodies. However, manual reading of the fluorescence signals is subjective and leads to high intra- and inter-laboratory variation, making standardized automated evaluation a desirable goal.
Automated interpretation of CLIFT has recently been incorporated into the EUROPattern system [3] (Figure 2). The software is able to recognize the organelles of the protozoan and evaluates the specific kinetoplast fluorescence rather than just dark-light classification, increasing the reliability of the evaluation. Results are classified as positive or negative, and include a titre designation based on the fluorescence intensity.
In a clinical study, automated and visual evaluation of C. luciliae IIF was compared using 569 consecutive sera submitted for routine anti-dsDNA screening and 100 sera from healthy blood donors. The automated system recognized all 73 of the anti-dsDNA positive samples identified by the visual evaluation. Moreover, 93% of the titre designations were concordant. The overall sensitivity of the system amounted to 100% with a high specificity of 97%. Compared to visual microscopy the overall accuracy was 97%.
ANCA
ANCA are important serological markers for diagnosis and differentiation of autoimmune vasculitides, especially granulomatosis with polyangiitis (GPA, formally known as Wegener’s granulomatosis), which is characterized by autoantibodies against proteinase 3 (PR3), and microscopic polyangiitis, which is typified by autoantibodies against myeloperoxidase (MPO). In addition, ANCA can be found in chronic inflammatory bowel diseases. ANCA are detected by IIF with monospecific confirmation using ELISA, immunoblot or IIF microdot assays.
The IIF substrates ethanol-fixed and formalin-fixed granulocytes are used to identify the typical ANCA staining patterns of anti-PR3 (cytoplasmic, cANCA) and anti-MPO (perinuclear, pANCA) antibodies. An additional substrate consisting of HEp-2 cells coated with granulocytes allows immediate differentiation between ANCA and ANA, while purified antigen microdots of PR3, MPO or glomerular basement membrane (GBM) antigen provide simultaneous monospecific antibody characterization. The different substrates are incubated and automatically evaluated in parallel as BIOCHIP mosaics, thus providing ANCA screening and confirmation in one step.
Evaluation software such as EUROPattern provides automated positive/negative discrimination of samples, as well as recognition of pANCA and cANCA patterns [1] (Figure 3). Further pattern constellations such as DNA-ANCA (atypical pANCA, xANCA), which can arise from antibodies against lactoferrin or other antigens, are also taken into account by the software. The automated system proposes a result based on the recognized cellular patterns and the results on the antigen microdots. An estimated titre with a confidence value is given.
Anti-neuronal antibodies
Neuronal cell-surface autoantibodies occur in autoimmune encephalitis and their detection can secure an early diagnosis, enabling immediate treatment which is critical for patient outcome. In recent years a considerable number of novel target antigens has been discovered, for example, glutamate receptors of type NMDA and AMPA, GABAB receptors, voltage-gated potassium channel-associated proteins LGI1 and CASPR2, DPPX and IgLON5.
Diagnostic tests for the new parameters are based on recombinant-cell (RC) IIF, in which transfected cells expressing the relevant antigen are used for monospecific antibody detection. This test method enables authentic presentation of the fragile membrane-associated surface antigens. Since many of the autoantibody markers are rare and do not always overlap, a multiparametric screening using BIOCHIP mosaics made up of different substrates is recommended. Results for RC-IIF assays can be evaluated automatically using a newly developed function of EUROPattern. The system automatically takes digital images of the substrates and provides a positive/negative classification.
The quality of the acquired images was assessed by comparing on-screen appraisal with visual microscopy using 753 incubations of numerous serum samples sent to a clinical immunology laboratory [4]. Ambiguous fluorescence signals detected at the microscope were excluded to avoid inter-reader deviations. The two evaluation strategies revealed a concordance of 100% with respect to positive/negative discrimination, confirming the high quality of the images.
Arbovirus antibodies
Immunofluorescence microscopy is also useful for infectious disease diagnostics. For example, infections with Zika virus, dengue virus and chikungunya virus are difficult to tell apart clinically as they manifest with similar symptoms and are endemic in much the same regions. Serological tests are an important diagnostic method, especially beyond the short viremic phase when direct detection is no longer effective. Viral antibodies can be detected by IIF using virus-infected cells. However, cross reactions between flavivirus antibodies can occur.
A BIOCHIP mosaic comprising substrates for Zika, dengue and chikungunya viruses enables parallel antibody determination, aiding clarification of cross reactivities and supporting differential diagnosis. The substrates can be evaluated semi-automatically using the digital image acquisition function of EUROPattern. In particular, inspection of the images side-by-side on the computer screen considerably facilitates the interpretation.
Fully automated immunofluorescence microscopy
Computer-aided fluorescence microscopy can be further standardized and facilitated through use of complementary hardware. The EUROPattern microscope (Figure 4) has been tailored to the requirements of immunofluorescence. Next to the high-precision optical system, it has a controlled LED, which maintains a constant light flux, ensuring highly reproducible results. The cLED has an extremely long life span without maintenance (over 50,000 hours) and low power consumption, ensuring cost-effectiveness for laboratories. The microscope is equipped with a slide magazine which can process up to 500 analyses in succession within 2.5 hours (18 seconds per field), correctly identifying the slides by means of matrix codes.
Results from the automated IIF evaluation can be viewed and validated directly at the computer screen, enabling a diagnosis to be established quickly and efficiently. The high-resolution images are sharply focused with the aid of a counterstain. The counterstain also serves to verify correct performance of the incubation. Negative results can be verified in batches, while positive samples can be individually checked and confirmed by the medical technologist. Results from different serum dilutions and substrates are consolidated into one report per patient, and new findings are compared with previous records. Final results can be signed electronically and forwarded at a click.
Perspectives
The need for standardization and automation in IIF is tremendous in all fields of autoimmune diagnostics. In particular, manual evaluation of results is time consuming and subjective. Automation platforms with harmonized software and hardware components have in recent years contributed enormously to the standardization and simplification of the evaluation process, especially for ANA, ANCA and CLIFT. Advanced software provides positive/negative classification, pattern recognition and titre designation at a quality equivalent to visual microscopy. The recording of tissue substrates, such as liver, kidney, stomach, esophagus, small intestine, heart and neuronal tissue, is also feasible. Future development will focus on the recognition of organ- and non-organ-specific autoantibodies on tissues, for example antibodies against mitochondria, epithelial membrane, epidermal basement membrane, desmosomes, heart muscle and neuronal antigens. The continued development of automated evaluation systems is anticipated to lead to even greater standardization of IIF and further reductions in workflow for diagnostic laboratories.
References
1. Krause et al. Lupus 2015: 24: 516-29
2. Voigt et al. Clin. Devel. Immunol. 2012: vol 2012, article ID 651058
3. Gerlach et al. J. Immunol. Res. 2015: vol 2015, article ID 742402
4. Fraune et al. Autoimmunity Reviews 15 (2016) 937-942
The author
Jacqueline Gosink, PhD
EUROIMMUN AG, Seekamp 31, 23560 Luebeck, Germany
E-mail: j.gosink@euroimmun.de