Lifespin launches commercial access to its metabolic profiler software and database for pharma and biobank services
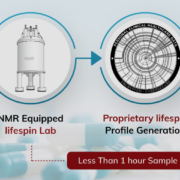
Lifespin, based in Regensburg, Germany, with offices in Boston, Massachusetts, has launched a new commercial service that, for the first time, will provide the pharmaceutical and biobanking industry with access to its proprietary metabolic database, as well as its advanced interpretive software to assist in various phases of drug research, development, and manufacturing.