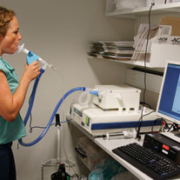
Diagnosing asthma in smokers
Asthma patients who smoke report more pronounced symptoms, an attenuated response to inhaled corticosteroids and more frequent attacks. Furthermore, diagnosing asthma in smokers can be difficult, as smoking impacts on the results of frequently used diagnostic and monitoring tools for asthma, including exhaled NO (eNO) and airway challenges.
by Dr Christian G. Westergaard, Professor Vibeke Backer and Dr Celeste Porsbjerg
Clinical background
Asthma is one of the most frequent chronic diseases worldwide, with an estimated global prevalence of 300 million people. The disease is characterised by respiratory symptoms, airway hyperresponsiveness (AHR) and bronchopulmonary inflammation. Asthma symptoms can be triggered by different agents, including exposure to allergens, physical activity and unspecific irritants such as air pollution, perfume, air humidity and tobacco smoke. The prevalence of asthma varies considerably between countries, however, in general, the prevalence has been increasing during recent decades, and ranges from a few percent to more than 15% in some countries [1].
It is estimated by the WHO that 1.25 billion people in the world are smokers. The global tobacco consumption was in 2000 estimated to be 15 billion cigarettes per day. This number is not expected to decrease until 2030, because the total number of smokers will become higher due to a larger world population [2].
Tobacco smoking has very damaging impacts on the asthmatic disease. Asthma patients who smoke report more pronounced symptoms, an attenuated response to inhaled corticosteroids, more frequent exacerbations and a higher mortality rate from asthma. Furthermore, these patients suffer from an accelerated decline in lung function, where both the highly reactive tobacco smoke and the chronic asthmatic inflammation in combination contribute to airway tissue destruction. Unfortunately, tobacco smoking is common among asthma patients, with a frequency of smokers at least as high as found in the rest of the population. In most countries, smokers constitute 15–40% of the population.
In the clinical setting, spotting the asthma patients among smokers can be challenging, due to the overlap of airway symptoms between true asthma and smoking-induced manifestations such as productive cough as well as breathlessness during exercise. In patients with a significant smoking history, an element of early chronic obstructive pulmonary disease (COPD) can also blur the clinical picture.
The diagnosing and monitoring of asthma has traditionally been based on the evaluation of symptoms in combination with spirometric measurements, which to date remain key elements in the clinical handling of asthma patients. However, as asthma is basically an inflammatory disease, many new diagnostic approaches focusing on airway inflammation have emerged, such as exhaled nitric oxide (eNO), sputum induction and airway challenges, of which the most recently approved is the mannitol test. All of these newer tests contribute to the understanding of the underlying pathophysiological mechanisms of the disease as well as expanding our diagnostic possibilities.
However, it appears that tobacco smoke may attenuate the clinical utility of many of the tests. In the following section, the focus will be on the effect of smoking on inflammation markers, AHR and spirometry, respectively.
Inflammation markers: eNO and induced sputum
Smoking has a considerable impact on the measurement of eNO. Several studies have reported a pronounced reduction of eNO in smokers compared to non-smokers, as much as 40–60% in current smokers [3]. Even passive smoking seems to have an effect on eNO values. Moreover, in a study from 2009 it was shown that eNO could only discriminate asthmatics from healthy controls in never-smokers, and not in either current or former smokers [4]. However, we have recently reported data from large sample, demonstrating that in adults with symptoms suggestive of asthma, eNO was equally good at differentiating between asthma and non-asthma, albeit with a lower cut-off for an abnormal eNO in smokers than in ex- and never smokers [5].
An eNO value of 17–22 ppb has been proposed for diagnosing asthma in current smokers [5, 6], supported by others who suggested 18 ppb as a cut-off for smokers without allergic rhinitis [6]. These similar cut-off values represent quite different sensitivity values, from about 40 to 100 %, when preserving a high specificity of at least 90%.
It would seem that eNO can also be applied in disease monitoring of the smokers when used in sequential measurements, because even in smokers, relative changes in eNO have been shown to reflect the dynamics of disease activity [7]. It has been demonstrated that, similar to non-smokers, a decrease in eNO of <20% precludes asthma control improvement, and that an increase in eNO of <30% is not associated with loss of control [7].
An important issue is the lack of knowledge regarding cut-off values for predicting steroid-response. In non-smokers, the effect of treatment with steroids has been found to be associated with airway eosinophilia, which again correlates well with eNO. Hence, a cut-off value for eNO predicting a sputum eosinophil count >3% and a high likelihood of a positive steroid response has been investigated. In smokers, this value was 28 ppb, ranging from 15 to 33 ppb, depending on atopy and high dose ICS usage [8], compared to 24 to 58 ppb in non-smokers. Such cut-off values are, however, not easy to determine, due to many factors of importance for the level of eNO, including atopy with rhinitis, life tobacco consumption and respiratory tract infections.
Several underlying mechanisms for the decreased eNO in smokers have been demonstrated, including increased arginase expression leading to reduced amounts of iNOS substrate, attenuated eosinophilic and enhanced neutrophilic inflammation as well as the impact on exogenous NO from cigarette smoke leading impairment of NO synthesis.
Another way of characterising the inflammation in the airway tissue is through sputum induction. This technique is rarely used in the diagnosis of asthma. In smoking asthmatics, it seems that the cell distribution is altered into a less eosinophilic and more neutrophilic direction. This may partially explain why smokers are less responsive to steroids.
Bronchial challenges: mannitol and methacholine
Another important approach in asthma diagnostics is measurement of airway hyperresponsiveness (AHR) using the mannitol challenge, which is an indirect bronchial provocation. In non-smokers, this test can be successfully applied for both diagnostic and monitoring purposes. It has been shown that the mannitol challenge is useful in confirming a diagnosis of asthma (specificity close to 100%), unfortunately, however, the sensitivity is considerably more moderate, around 60% [9] and thereby lower than that of the methacholine challenge [9]. Being a relative recent invention, the diagnostic properties of the mannitol test have not yet been evaluated in a smoking asthmatic population. However, in non-asthmatic smokers, a study has indicated that as much as one quarter of the subjects expressed a positive mannitol test [10]. Thus, until investigated properly in smoking asthma patients, the mannitol challenge test should be interpreted with caution and be accompanied with other tests in order to account for false positives.
AHR can also be assessed through direct challenges such as inhaled methacholine. The higher sensitivity for the methacholine test (69%) compared to the mannitol test is, unfortunately, not accompanied by an equivalently higher specificity, which has been reported to be 80% [9]. In non-asthmatics, previous studies have indicated increased AHR to methacholine in smokers. But as is the case with the mannitol test, the diagnostic properties of the methacholine test have not yet been investigated in a smoking asthmatic population, which is surprising considering that the test has been applied for decades. However, a few studies have documented that smoking does appear to affect AHR to both direct and indirect challenges. In COPD patients, it has been shown that one year of smoking cessation is associated with improvement in AHR to methacholine as well as to AMP; this finding has been supported later in a study primarily of healthy subjects, but also a few asthma patients.
Spirometry with reversibility test
Increased bronchial muscular tonus is a key feature in persistent asthma, which is the reason that measurements of lung function, including the reversibility test, have been widely used in asthma diagnostics and monitoring for decades. For some smoking asthma patients, this will continue to be a corner stone in confirming the diagnosis, but in smoking asthmatic subjects with a baseline normal FEV1 or patients with very severe asthma and hence attenuated airway compliance, reversibility testing may not be the best diagnostic test. Many studies of asthma and COPD patients have shown improvement in FEV1 after smoking cessation, indicating an airway narrowing effect of tobacco smoke. However, a study of 134 asthma patients with airway reversibility showed no difference in baseline FEV1 between smokers and non-smokers, and the salbutamol reversibility was similar [11]. This latter finding has also been confirmed in a few other studies, but, in general, our knowledge of the effect of smoking on the β2-agonist reversibility of airway resistance is sparse.
Conclusion
Smoking affects the results of most of the different clinical asthma tests available, and test results should be interpreted with smoking status in mind. Clinicians should be aware of potential limitations of each test, especially eNO, which decreases in smokers but remains useful, and the mannitol test, which may give false positive in smokers. It remains crucial to obtain an explorative anamnestic interview, involving clarification of symptom triggers, seasonal variation, presence of wheezing, concomitant rhinitis, night symptoms, familiar dispositions, symptom debut, allergies and of course, smoking history.
References
1. Masoli M, Fabian D, Holt S, Beasley R. Allergy. 2004; 59(5): 469-478.
2. Annual global cigarette consumption. http://www.who.int/tobacco/en/atlas8.pdf
3. Alving K, Malinovschi A. Eur Respir Mon 2010; 49: 1-31.
4. Malinovschi A, Janson C, Högman M, Rolla G, Torén K, Norbäck D, Olin AC. Allergy. 2009; 64(1): 55-61.
5. Malinovschi A, Backer V, Harving H, Porsbjerg C. Respir Med. 2012; 106(6): 794–801.
6. Matsunaga K, Hirano T, Akamatsu K, Koarai A, Sugiura H, Minakata Y, Ichinose M. Allergol Int. 2011; 60(3): 331-337.
7. Michils A, Louis R, Peché R, Baldassarre S, Van Muylem A. Eur Respir J. 2009; 33(6): 1295–301.
8. Schleich FN, Seidel L, Sele J, Manise M, Quaedvlieg V, Michils A, Louis R. Thorax. 2010; 65(12): 1039-1044.
9. Sverrild A, Porsbjerg C, Thomsen SF, Backer V. J Allergy Clin Immunol. 2010; 126(5): 952–958.
10. Stolz D, Anderson SD, Gysin C, Miedinger D, Surber C, Tamm M, Leuppi JD. Respir Med. 2007; 101(7): 1470-1476.
11. Chaudhuri R, McSharry C, McCoard A, Livingston E, Hothersall E, Spears M, Lafferty J, Thomson NC. Allergy. 2008; 63(1): 132-135.
The authors
Christian G. Westergaard MD*,
Vibeke Backer MD, DMSc and
Celeste Porsbjerg MD, PhD
Bispebjerg Hospital
Respiratory Research Unit
Bispebjerg Bakke 23, Entrance 66
DK-2400 Copenhagen NV, Denmark
*Corresponding author:
e-mail: cgwestergaard@hotmail.com