Systematic multiplex PCR for the diagnosis of infectious gastroenteritis
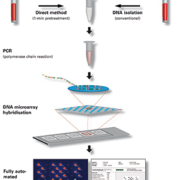
Current methods for the detection of gastroenteric pathogens are insensitive, slow, and laborious. Testing directed at specific organisms misses important infections. Systematically testing all fecal samples using multiplex PCR for common viral, bacterial and parasitic pathogens allows laboratories to increase diagnostic yield, improve workflow, reduce waste and turn-around times.
by Dr Gary McAuliffe
Introduction
Within a single diagnostic laboratory, multiple methods are used to detect gastroenteric pathogens. Selective agar plates differentiate bacterial pathogens, whereas immunoassays are used to detect viruses and parasites such as Giardia lamblia and Cryptosporidium spp.. Laboratories perform microscopy with special stains for the detection of Entamoeba histolytica and Dientamoeba fragilis. PCR has generally been restricted to the detection of norovirus, but many studies have demonstrated its potential for the detection of other enteric pathogens.
Multiplex PCR (M-PCR) panels have been shown to enhance detection of gastroenteric organisms. These panels combine several enteric pathogen targets in one or more PCR reaction vessel(s). O’ Leary et al. demonstrated 100% sensitivity compared with culture for the detection of four bacterial pathogens [1]. Wolffs et al. used a panel targeting seven viral pathogens and detected a pathogen in 97% of samples compared with 49% by conventional methods [2]. Stark et al. showed 100% sensitivity and specificity of a M-PCR panel targeting four parasites, which also reliably differentiates E. histolytica from non-pathogenic E. dispar and E. moshkovskii [3]. de Boer et al. successfully replaced bacterial culture at their institution with a molecular screening approach targeting four bacteria and G. lamblia [4].
Several commercial M-PCR fecal panels are available [Table 1]. These may be directed against parasites, viruses or bacteria, or contain targets from all three groups of organisms, allowing systematic testing of stool samples for all common gastrointestinal pathogens.
In the author’s study, 1758 samples from community and hospital patients were tested using the Fast-Track Diagnostics bacterial, viral and parasite M-PCR panels. Pathogens were detected in 30% of samples by this systematic M-PCR approach compared with 18% using conventional testing as directed by clinician request [5][Table 2].
Advantages of a systematic M-PCR approach
Studies have demonstrated enhanced detection of entero-hemorrhagic E. coli (EHEC), Clostridium difficile, G. lamblia, E. histolytica, norovirus, adenovirus and rotavirus by PCR compared with immunoassays and other conventional assays [2–4, 6, 7]. Bacterial PCR has generally performed comparably with culture, with the exception of Salmonella spp., where reduced detection by PCR has been demonstrated in several, but not all, studies [4, 5, 8]. An advantage of increased sensitivity is that smaller amounts of feces are required for testing, and multiple samples are not required for the detection of common parasites. Clinicians should be aware that organisms such as adenovirus and norovirus can be shed in feces for several weeks following infection and that PCR detects low levels of these organisms which may not be clinically relevant. Selective bacterial media lack specificity, requiring time and further testing to discount commensal organisms. E. histolytica cysts cannot be reliably differentiated from other members of the Entamoeba complex by microscopy and staining. M-PCR generally overcomes these issues, though specificity depends on the target sequence chosen. For adenovirus, some panels target the hexon gene which does not differentiate between enteric and non-enteric serotypes, whereas others target sequences specific to enteric sub-types.
Currently laboratories restrict their testing to a limited range of pathogens for which they have sensitive and affordable assays available. A number of organisms that laboratories do not commonly test for, such as astrovirus, sapovirus, Vibrio spp., and non-O157 EHEC, can be missed. M-PCR allows a wider range of organisms to be targeted. In the author’s study astrovirus was found to be the second most common cause of gastroenteritis, and non-O157 EHECs were detected in sixteen samples by targeting the stx genes. Our laboratories did not have assays for these organisms prior to the study. Laboratories may design their own panels to reflect locally and internationally important pathogens, or buy commercial panels which reflect these requirements. Rare or imported infections not targeted by the panels will not be detected, and laboratories need to decide when additional tests are required. Up to five targets may be tested in a single real-time PCR reaction vessel; therefore increasing the number of targets reduces the number of samples on a given PCR amplification run. The xTAG system (Luminex Corp.) overcomes this limitation by post-amplification analysis using microspheres with up to 100 differing spectra. Eleven targets are currently included in the xTAG gastroenteritis panel within a single PCR reaction vessel [9]. TaqMan array cards (Life technologies) also overcome this restriction by allowing simultaneous real-time PCR in 384 PCR reaction wells. This platform allows up to eight samples to be run in parallel [10].
Fecal samples are usually tested for a limited range of viruses, bacteria, or parasites dependent upon clinician request and laboratory algorithms. Studies have shown that important pathogens such as G. lamblia and EHEC are missed using this approach. Testing for enteric viruses is not widely employed outside hospitals despite their prevalence. The M-PCR approach tests every sample for every target in the panels. It is less reliant on clinicians’ knowledge of the organisms that the patient has likely been exposed to, or those that may be causing the patients clinical syndrome. This systematic approach accounted for the majority of the increased diagnostic yield in the author’s study.
In our laboratory it can take 3–4 days for the identification of Salmonella spp. by culture. Time to generation of results is significantly reduced with the M-PCR approach. In the author’s study, samples collected were batch tested the following day, giving results for eleven pathogens simultaneously within 24 hours of collection. Hands-on time is also reduced. The preparation and reading of a trichrome stain can take up to 40 minutes by an experienced operator, whereas the hands-on time for testing a sample by PCR is a quarter of this.
The workflow created by the systematic M-PCR approach fits well with current laboratory systems; testing is performed in a single pathway rather than by several laboratory departments. Conventional fecal testing employs multiple diagnostic kits and selective media, some of which come with significant cost, short expiry times, and extensive quality control requirements. In our laboratory stool samples for bacterial culture are inoculated onto seven agar plates which need to be stored, processed, and incubated, generating a significant amount of waste. O’ Leary et al. reported that M-PCR significantly reduced this wastage [1]. M-PCR does not obviate the need for bacterial culture as it does not offer an antibiogram, but it allows focused testing of samples found to be positive for these bacterial targets.
Pathogens such as Shigella spp, G. lamblia and D. fragilis are labile in stool. Delays may occur in transportation, inoculation or examination which can compromise yield. For M-PCR less initial processing is required. Specimens are inoculated into tubes containing Stool Transport and Recovery buffer (STAR) (Roche Diagnostics) which binds inhibitors, and stabilizes nucleic acids for later processing. PCR is also able to detect organisms rendered non-viable by inadequate transport, or the use of antibiotics. Feces contains high levels of bilirubin and bile salts, which can lead to inhibition of PCR amplification. Inoculating samples into STAR buffer on arrival, and using extraction methods such as the EasyMag platform (Biomerieux) can help overcome this issue. In the author’s study 1.7% of samples exhibited inhibition, but all gave adequate internal control amplification following dilution and repeat testing.
Cost is a major factor preventing systematic testing of fecal samples by conventional techniques in diagnostic laboratories. The costs of PCR are reducing relative to conventional tests, and batch testing of samples using the systematic M-PCR approach is becoming a viable option. In the author’s study testing a sample for bacteria, viruses and parasites was significantly cheaper by M-PCR than using equivalent conventional tests [NZ$152 (£75) versus NZ$280 (£140)]. Laboratories have successfully reported replacing their conventional methods with a molecular screening approach for bacteria [1, 4] or viruses [2]. Where the cost of replacing all traditional diagnostics with systematic testing may currently remain restrictive, laboratories may choose to replace testing for organism groups, e.g. viruses, and institute systematic testing at a later date.
Conclusion
Current conventional methods for the detection of enteric pathogens are labour intensive, insensitive, slow, and applied piecemeal to submitted fecal samples. Testing stool samples using multiplex PCR panels which simultaneously detect all common bacteria, viruses and parasites increases the detection of gastroenteric pathogens. This approach improves turn-around time, workflow, reduces labour and waste. Costs are reducing, making systematic M-PCR testing an attractive alternative to currently used techniques.
References
1. O’Leary J, et al. Comparison of the EntericBio multiplex PCR system with routine culture for detection of bacterial enteric pathogens. J Clin Microbiol. 2009; 47: 3449–3453.
2. Wolffs PF, et al. Replacing traditional diagnostics of fecal viral pathogens by a comprehensive panel of real-time PCRs. J Clin Microbiol. 2011; 49: 1926–1931.
3. Stark D, et al. Evaluation of multiplex tandem real-time PCR for detection of Cryptosporidium spp., Dientamoeba fragilis, Entamoeba histolytica, and Giardia intestinalis in clinical stool samples. J Clin Microbiol. 2011; 49: 257–262.
4. de Boer RF, et al. Improved detection of five major gastrointestinal pathogens by use of a molecular screening approach. J Clin Microbiol. 2010; 48: 4140–4146.
5. McAuliffe GN, et al. Systematic application of multiplex PCR enhances the detection of bacteria, parasites, and viruses in stool samples. J Infect. 2013; 67(2): 122–129.
6. Costantini V, et al. Diagnostic accuracy and analytical sensitivity of IDEIA norovirus assay for routine screening of human norovirus. J Clin Microbiol. 2010; 48: 2770–2778.
7. Luna RA, et al. Rapid stool-based diagnosis of Clostridium difficile infection by real-time PCR in a children’s hospital. J Clin Microbiol. 2011; 49: 851–857.
8. Cunningham SA, et al. Three-hour molecular detection of Campylobacter, Salmonella, Yersinia, and Shigella species in feces with accuracy as high as that of culture. J Clin Microbiol. 2010; 48:2929–2933.
9. Coste JF, et al. Microbiological diagnosis of severe diarrhea in kidney transplant recipients by use of multiplex PCR assays. J Clin Microbiol. 2013; 51(6): 1841–1849.
10. Liu J, et al. A laboratory developed TaqMan array card for simultaneous detection of nineteen enteropathogens. J Clin Microbiol. 2013; 51(2): 472–480.
The author
Gary McAuliffe MBBS
Microbiology Department, LabPlus Laboratory, Auckland, New Zealand
E-mail: GMcAuliffe@adhb.govt.nz